Tandem Mass Spectrometry-Based Peptide Sequence Analysis Service
-
Original LC-MS/MS data files;
-
Sequence interpretation and modification information tables (Excel format);
-
Fragment ion spectra and peak plot files (TIFF or PNG format);
-
Comprehensive analysis report (PDF format) including experimental conditions and result descriptions.
MtoZ Biolabs has launched the tandem mass spectrometry-based peptide sequence analysis service which enables highly sensitive and precise analysis of peptide sequences. This service is widely applied in peptide drug development, protein structure research, and quality control, providing reliable data support for peptide sequence elucidation and functional studies.
Principle of Tandem Mass Spectrometry
Tandem mass spectrometry (MS/MS) is a high-resolution analytical technique that determines molecular structures through multi-stage mass spectrometric analysis. Its principle involves selecting target ions (parent ions) in the first mass analyzer, inducing fragmentation through collision, and then detecting the resulting fragment ions in the second mass analyzer to deduce molecular structure information. In peptide sequencing, MS/MS accurately identifies amino acid sequences, modification sites, and molecular weight variations. This technology offers high sensitivity, resolution, and throughput, making it widely used in proteomics research, peptide drug discovery, and biomarker identification.
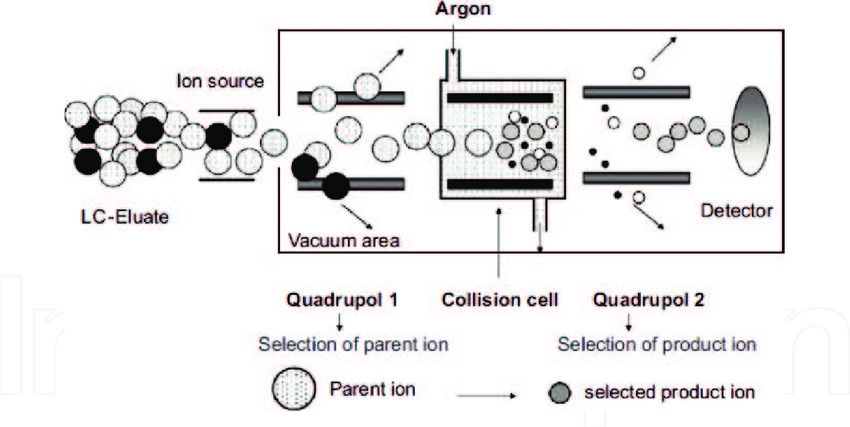
Baykalir, Y. et al. New Insights into Theriogenology, 2018.
Figure 1. Principle of Tandem Mass Spectrometry (MS/MS).
Tandem Mass Spectrometry-Based Peptide Sequence Analysis Service at MtoZ Biolabs
1. Sequence Verification
Accurately analyzes the amino acid sequence of peptides using high-resolution tandem mass spectrometry to verify consistency with the target sequence.
2. Modification Analysis
Detects and locates post-translational modifications in peptides, such as phosphorylation, methylation, and acetylation, revealing their impact on biological function.
3. Molecular Weight Determination
Precisely determines peptide molecular weight using high-sensitivity mass spectrometry to confirm the correctness of synthesized or expressed products.
4. Fragment Ion Analysis
Interprets MS/MS fragmentation spectra to analyze b-ion and y-ion series, assisting in reconstructing complete sequences and enhancing identification confidence.
Workflow of Tandem Mass Spectrometry-Based Peptide Sequence Analysis Service
1. Sample Processing and Separation
Peptide samples are purified, desalted, and separated by liquid chromatography to ensure optimal detection sensitivity and resolution.
2. Parent Ion Selection and Fragmentation
Target parent ions are selected in the first mass spectrometer and fragmented through collision-induced dissociation to generate characteristic fragment ions.
3. Tandem Mass Spectrometry Detection
High-resolution MS/MS is used to obtain fragment ion spectra, providing the data foundation for sequence interpretation.
4. Data Analysis and Sequence Deduction
Fragment ion patterns are analyzed to determine amino acid sequences, modification sites, and mutation features.
5. Result Integration and Report Generation
A comprehensive analysis report is generated, including spectra, sequence information, and modification annotations.
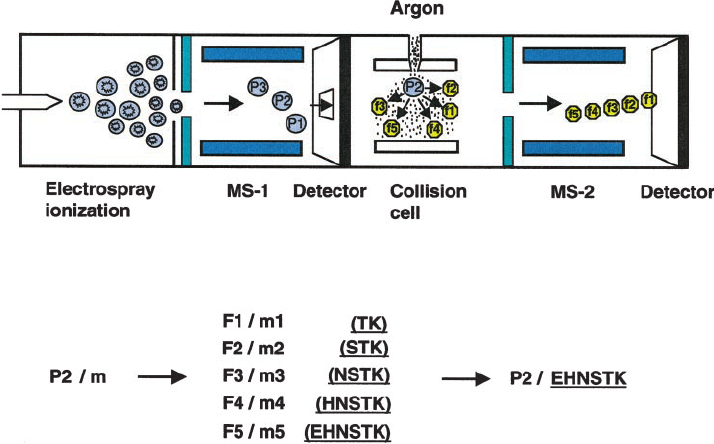
Li, J X. et al. Plant Physiology, 2000.
Figure 2. Schematic of Peptide Sequencing by Tandem Mass Spectrometry.
Why Choose MtoZ Biolabs?
✅ High-Resolution Detection: Tandem mass spectrometry enables high-precision analysis of amino acid sequences and modifications.
✅ High-Sensitivity Analysis: Capable of detecting low-abundance peptides and identifying trace modification sites.
✅ Fast Result Delivery: Efficient data processing and automated analysis significantly shorten turnaround time.
✅ Professional Technical Support: Personalized solutions provided by an experienced mass spectrometry expert team.
✅ One-Stop Service: Covers the entire process from sample preparation and detection to data analysis and report generation.
Applications of Tandem Mass Spectrometry-Based Peptide Sequence Analysis Service
1. Protein Structure Research
Reveals the origin and domain features of protein fragments through sequence analysis, providing a basis for structural studies.
2. Modification and Mutation Detection
Identifies post-translational modifications and mutation sites in peptides to study their effects on function and stability.
3. Biomarker Screening
Analyzes sequence variations in disease-associated peptides to assist in the discovery of diagnostic and prognostic biomarkers.
4. Metabolite Analysis
Detects endogenous peptide sequences in cells or tissues to elucidate metabolic pathways and regulatory mechanisms.
5. Signaling Pathway Analysis
Identifies peptides involved in signal transduction to explore their roles in cellular communication and regulation.
Deliverables
1. Experimental Procedures
2. Relevant Mass Spectrometry Parameters
3. Detailed Information on MS/MS Peptide Sequences
4. Mass Spectrometry Spectra
5. Raw Data Files
FAQ
Q1: What types of samples are suitable?
A1: The tandem mass spectrometry-based peptide sequence analysis service is suitable for various types of peptide samples, including synthetic, recombinant, and naturally derived peptides. To ensure the accuracy and sensitivity of tandem mass spectrometry detection, samples should have high purity, be free from salts, buffers, and surfactant residues, and remain stable without degradation.
Q2: What is the service general workflow?
A2:
Q3: What data formats are provided?
A3: The service delivers standardized, multi-format data results to facilitate scientific analysis and record keeping. Regular deliverables include:
If special analytical requirements exist, data formats can be customized according to project specifications.
Q4: How should I prepare the samples?
A4: To ensure detection quality and result reproducibility, it is recommended that sample preparation follows these guidelines:
(1) Purity requirements: Avoid high salt, buffer, and other impurities.
(2) Storage conditions: Store at -20℃ for short term and -80℃ for long term.
(3) Transportation: Use cold chain or dry ice shipping to ensure sealed, leak-proof containers.
(4) Additional information: Provide sample source, concentration, and intended analysis goals to optimize testing parameters.
For more information, please refer to Sample Submission Guidelines for Proteomics, Sample Submission Guidelines for Metabolomics.
Start Your Project with MtoZ Biolabs
Contact us to discuss your experimental design or request a quote. Whether you are exploring peptide sequence characteristics or studying modification patterns and structural differences, MtoZ Biolabs can provide you with precise and efficient tandem mass spectrometry analysis support.
How to order?







