Resources
Proteomics Databases
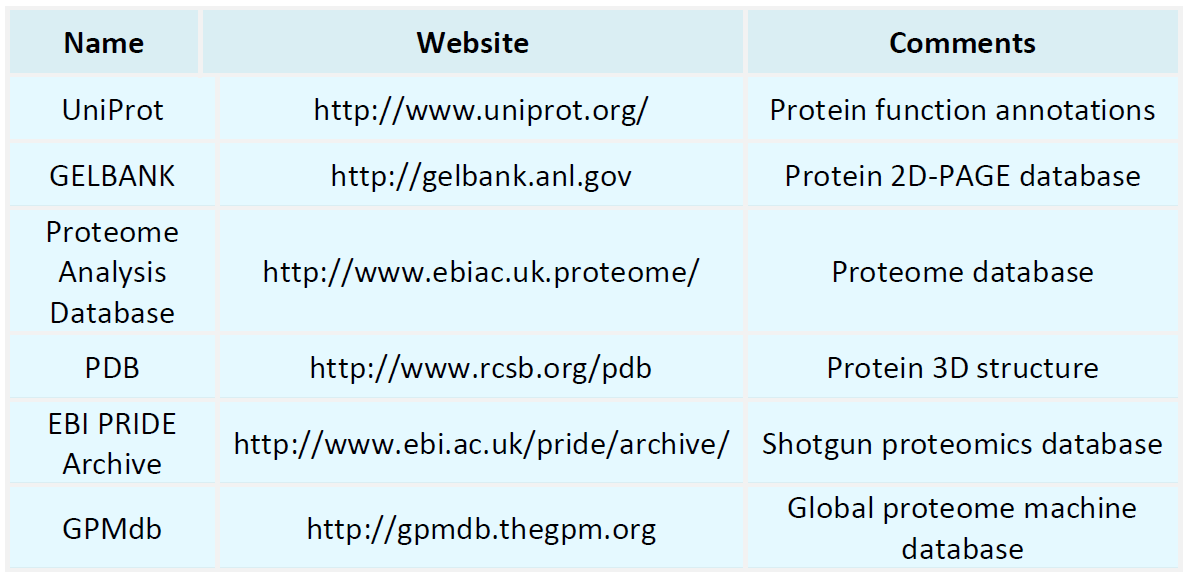
Metabolomics Databases
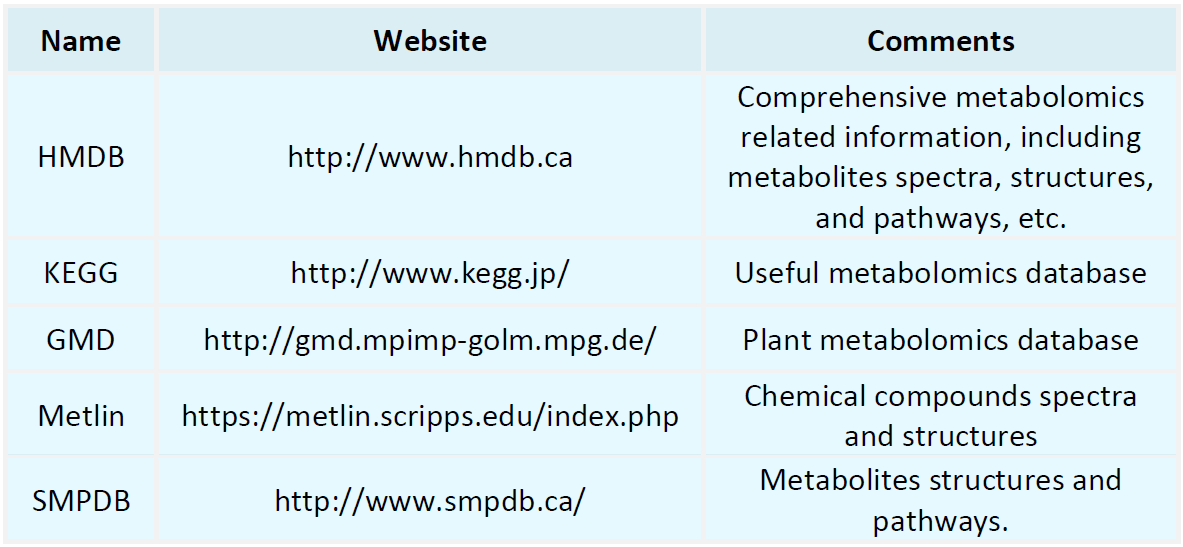
-
• How to Perform Quantitative Exosome Proteomics Using TMT/DIA?
Exosomes are cell-secreted vesicles with diameters ranging from 30 to 150 nm, widely distributed in blood, urine, and other body fluids. Rich in proteins, RNA, and lipids, exosomes serve as critical mediators of intercellular signaling and play significant roles in disease diagnosis, drug target identification, and therapeutic research. With advances in mass spectrometry, high-throughput and precise protein quantification methods have become essential tools for exosome research. Among these, Tandem Ma......
-
• What Are the Common Methods for Histone Propionylation Analysis?
In the rapidly evolving field of epigenetics, post-translational modifications of histones have emerged as a central focus in life sciences research. Among these, histone propionylation (Kpr), an emerging lysine acylation modification, has attracted considerable attention due to its potential roles in metabolic regulation and gene expression. Histone propionylation occurs on lysine residues and is chemically similar to acetylation, but with an additional carbon atom (C3). Even this minor structural di......
-
• What Enzymes Regulate Histone 2-Hydroxyisobutyrylation?
Over the past decade, epigenetics has advanced rapidly. In particular, the discovery of histone post-translational modifications (PTMs) has provided new insights into the regulation of gene expression. Beyond classical acetylation and methylation, histone 2-hydroxyisobutyrylation (2-hydroxyisobutyrylation, Khib) has emerged as a growing focus of scientific research. What is Histone 2-Hydroxyisobutyrylation? Histones are the core proteins of chromatin, and their N-terminal tails are enriched in lysine......
-
• What Does Histone Kbhb Reveal About Epigenetic Regulation?
In epigenetics, histone modifications constitute a central mechanism for the regulation of gene expression. Histone β-hydroxybutyrylation (Kbhb), a recently identified modification, plays a critical role in modulating chromatin structure and gene transcription. Kbhb refers to the addition of a β-hydroxybutyryl group to lysine residues, and its abundance is strongly influenced by cellular metabolic states, particularly increasing under conditions such as fasting, low-carbohydrate diets, or elevated ket......
-
A practical guide to quantitative protein profiling using LC-MS/MS, covering sample preparation, label-free and DIA strategies, data analysis, applications, and method selection.
-
• How Does β-Hydroxybutyryl-CoA Drive Histone Kbhb Formation?
Within cells, metabolism and gene expression are tightly interconnected. In recent years, histone modifications have been shown to serve not only as central mechanisms of epigenetic regulation but also as recorders of cellular metabolic states. Among these, histone β-hydroxybutyrylation (lysine β-hydroxybutyrylation, abbreviated as Kbhb) has emerged as a novel epigenetic mark that has attracted widespread attention. How, then, does the metabolic intermediate β-hydroxybutyryl-CoA drive the formation of......
-
• Common Challenges in LC-MS/MS Protein Identification and How to Solve Them
In proteomics research, LC-MS/MS (liquid chromatography-tandem mass spectrometry) has emerged as a central analytical technique for protein identification. Owing to its high sensitivity and high-throughput capacity, LC-MS/MS enables large-scale identification and quantification of proteins in complex biological samples. However, throughout experimental workflows and downstream data analysis, researchers frequently encounter challenges such as low protein identification yield, poor reproducibility, and......
-
• How to Prepare Samples for Histone Butyrylation Analysis?
Histones, as fundamental components of chromatin, are extensively involved in the regulation of gene expression, cell cycle progression, and DNA repair through diverse post-translational modifications. Among these, histone butyrylation represents an emerging modification structurally related to acetylation, with the capacity to influence chromatin architecture and transcriptional activity. Consequently, accurate identification and quantification of histone butyrylation have become critical in epigenet......
-
• How to Choose the Right Database for Proteomics Protein Identification?
In modern proteomics research, protein identification constitutes a key step. Advances in mass spectrometry have enabled researchers to acquire extensive peptide datasets from complex biological samples. However, peptides represent only fragments of information, and accurately mapping these fragments to specific proteins relies on high-quality protein databases. The choice of an appropriate database not only affects identification accuracy but also directly impacts the reliability of subsequent data a......
-
• What Is Histone Crotonylation Analysis?
Histone post-translational modifications (PTMs) represent a key mechanism regulating chromatin structure and gene expression. In recent years, with advances in mass spectrometry technologies, histone crotonylation (Kcr), a novel acylation modification, has attracted increasing attention. As a short-chain unsaturated acyl modification occurring on lysine residues, crotonylation is structurally distinct from classical acetylation and exhibits unique biological functions in transcriptional regulation, ce......
How to order?







