PROTAC Drug Off-Target Protein Assessment Service
Proteolysis targeting chimera (PROTAC) drugs offer a novel approach to disease treatment by leveraging the body's innate protein degradation pathways. In contrast to traditional small-molecule inhibitors, PROTAC drugs promote the degradation of specific proteins rather than directly inhibiting their function, thus mitigating or eliminating their role in disease progression.
PROTAC drugs have three main components: a ligand that binds to the protein of interest (POI), a ligand that binds to an E3 ubiquitin ligase, and a linker connecting the two ligands. Upon cell entry, PROTAC molecules bind to both the target protein and E3 ligase, forming a ternary complex. This leads to ubiquitination of the target protein, which is subsequently recognized and degraded by the proteasome.
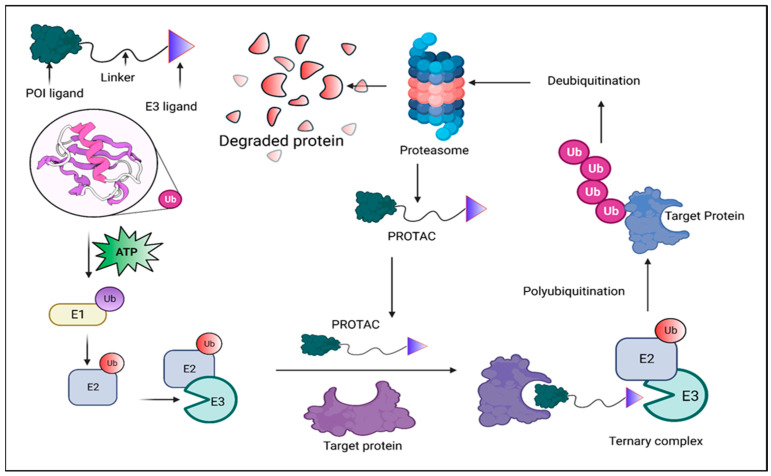
Figure 1. Mechanism of PROTAC Degradation [1]
The off-target effects of PROTAC drugs refer to the unintended degradation of non-target proteins, which can cause adverse side effects or toxicity. This occurs because PROTACs recruit E3 ubiquitin ligases to tag the target proteins for degradation. However, if the E3 ligase binds to unintended proteins, it may cause their unintended degradation.
Advantages and Limitations of PROTAC Drugs as Therapeutic Agents
1. Advantages
(1) Targeting "Undruggable" Proteins: PROTAC technology can degrade proteins that are challenging to target with traditional small-molecule drugs, expanding the potential range of druggable targets.
(2) Catalytic Inhibition: As catalytic agents, PROTACs can theoretically circulate within cells, reducing the necessary drug dose and potentially improving safety with fewer side effects.
(3) Overcoming Drug Resistance: PROTAC can help overcome resistance arising from genetic mutations by degrading pathogenic proteins.
(4) New Strategy for Drug Development: PROTAC provides a novel approach to drug development, particularly in cancer, immune disorders, viral infections, and neurodegenerative diseases.
(5) Functional Protein Knockout: PROTAC serves as a valuable tool for functional protein depletion, supplementing genetic knockout techniques and aiding fundamental biological research.
2. Limitations
(1) Development Challenges: PROTAC design and optimization are more complex than traditional small-molecule drugs. Current high-throughput screening methods are inadequate, and design principles are largely unknown, leaving much of PROTAC discovery empirical.
(2) Oral Bioavailability: PROTACs generally have high molecular weights and large polar surface areas, reducing membrane permeability and oral bioavailability.
(3) Dosage and Manufacturing Costs: Although PROTAC theoretically enables low-dose therapy, clinical trial doses are currently relatively high, leading to increased manufacturing costs.
(4) Toxicology and Side Effects: As PROTAC works by degrading proteins, on- and off-target effects could occur. Its toxicological profile is not well characterized, and side effects remain largely unknown.
(5) Selectivity and Specificity: Ensuring the selective degradation of target proteins without affecting non-targets is a challenge that requires careful development of highly selective PROTAC molecules.
(6) Drug Resistance: PROTACs utilizing different E3 ligases may encounter resistance issues that need further study.
Strategies to Overcome Limitations Associated with PROTAC Drugs
1. Molecular Design Optimization
Improve the cell permeability and oral bioavailability of PROTACs through chemical modifications, reducing polar interactions, and enhancing pharmacokinetic (PK) properties.
2. Expand E3 Ubiquitin Ligase Applications
Explore additional E3 ligases to diversify PROTAC designs and broaden their application.
3. Leverage Nanotechnology
Enhance water solubility and cellular permeability of PROTACs with nanoparticles and other delivery systems for selective accumulation in tumor tissues.
4. Expand Clinical Treatment Range
Apply PROTAC technology beyond oncology to other diseases like neurodegenerative disorders and inflammation/immunology to maximize its therapeutic potential.
5. Develop Novel PROTAC Models
Besides traditional small-molecule PROTACs, develop novel bioPROTACs and hybrid PROTACs to overcome conventional limitations.
6. Precision Targeting Delivery Systems
Implement smart premedication and drugtamer-PROTAC conjugates using tumor microenvironment-sensitive linkers to enhance tumor targeting.
7. Establish Clinical Research and Evaluation Systems
Optimize screening, druggability, and pharmacology systems to better assess the bioactivity and safety of PROTACs.
8. Explore Novel Therapeutic Targets
Find optimal targets for protein degradation, especially those "undruggable" targets that are difficult to target with conventional small molecule drugs.
9. Overcome Drug Resistance
Develop PROTACs that degrade pathogenic proteins causing resistance due to overexpression or mutation.
10. Clinical Translation Strategy
To facilitate clinical translation, develop effective delivery strategies to improve PK properties and clinical potential.
Considerations for Preclinical Safety Evaluation of PROTAC Drugs
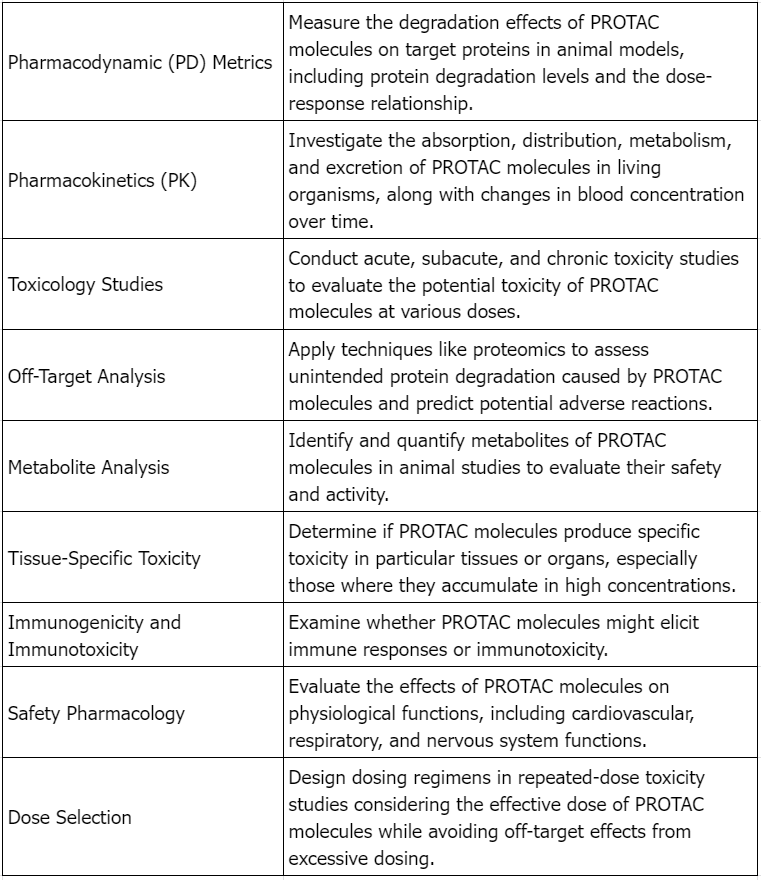
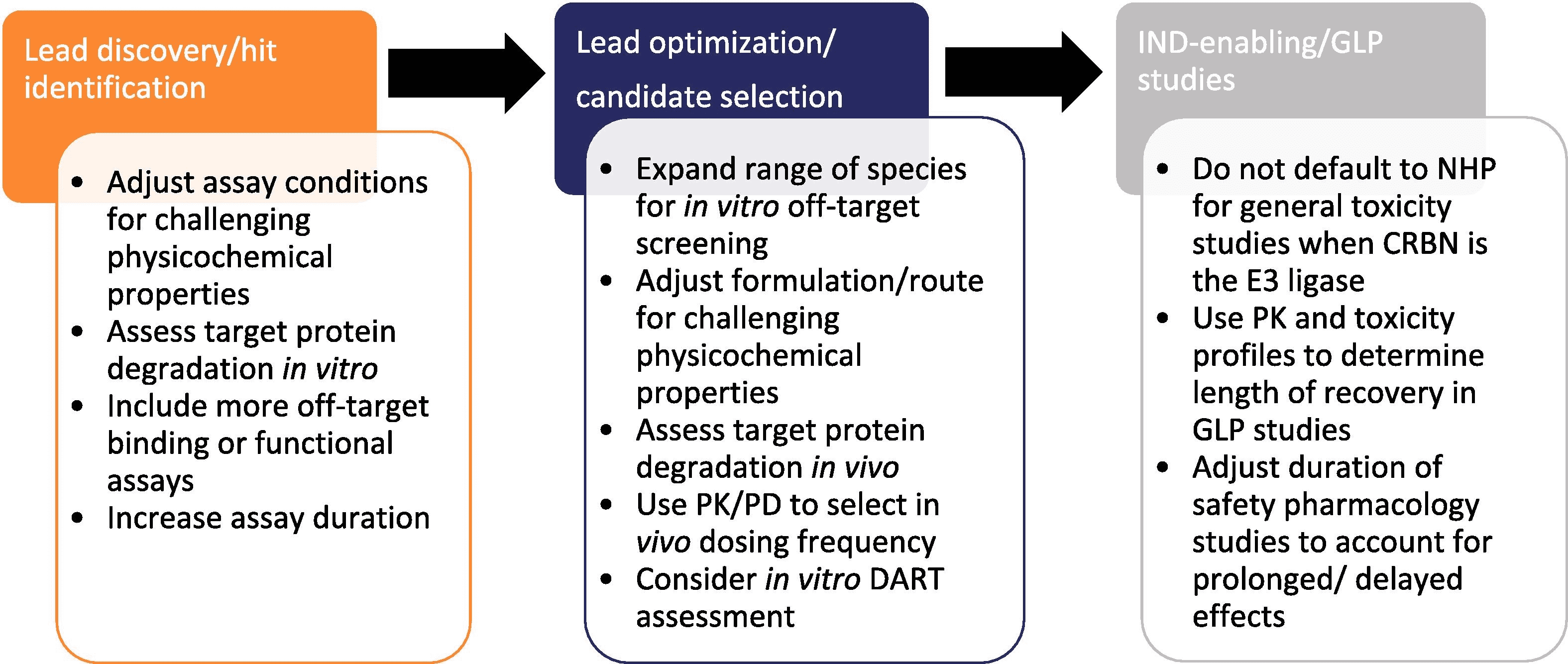
Figure 2. Considerations for Preclinical Safety Evaluation of Targeted Protein Degradation Agents [2]
PROTAC Drugs Analysis Solutions
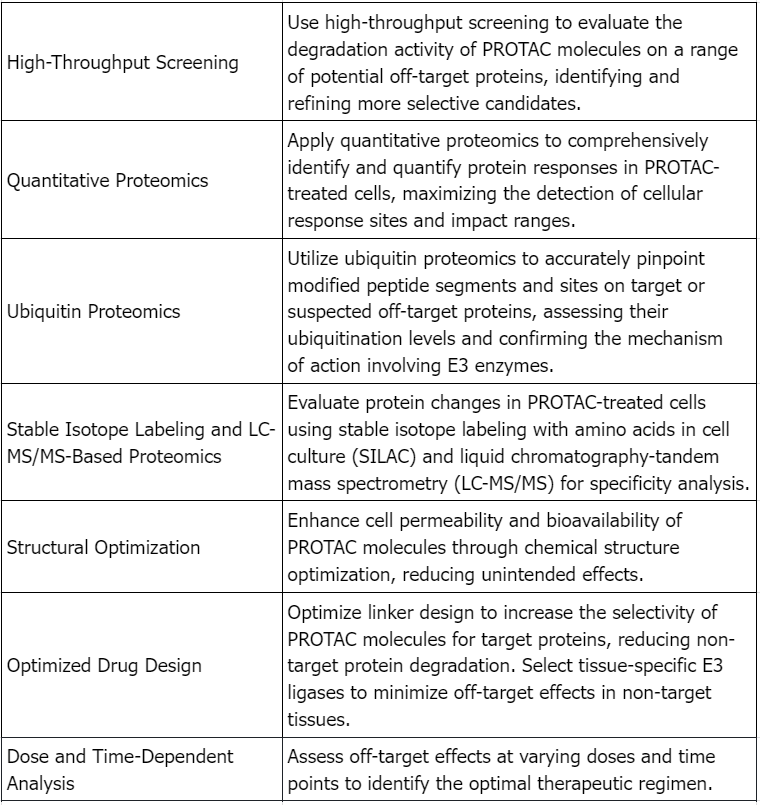
Service Advantages
1. Professional and Robust Services Covering a Wide Range of PROTAC Drugs Off-Target Analysis Testing
2. High Cost-Effectiveness, Short Testing Cycles, and Reliable, Comprehensive Test Reports
3. Equipped with a Variety of High-Resolution MS and other Major Equipment
Sample Results
1. A Proteomics Platform for Identifying Off-Target Proteins Related to Therapeutic Strategies Involving Protein Degradation or Gene Silencing
New therapeutic modalities like PROTAC and RNAi can inadvertently impact the abundance of endogenous proteins. The currently available secondary in vitro assays for assessing small molecules' off-target binding or activity cannot fully evaluate PROTAC off-target effects and are also unsuitable for RNAi. To fill this gap, researchers created a proteomics-based platform to comprehensively assess off-target protein abundance. They first selected off-target proteins using genetic and pharmacological evidence, which resulted in a set of 2,813 proteins called the "selected off-target proteome" (SOTP). An iterative algorithm then identified four human cell lines from a pool of 932, which, according to transcriptomic data, together express ~80% of the SOTP. Mass spectrometry then quantified intracellular and extracellular proteins in these cell lines. Of over 10,000 quantifiable proteins, 1,828 were part of the predefined SOTP. The SOTP was designed to be easily modified or expanded, owing to the rational selection process developed and the label free LC-MS/MS approach chosen. This platform's flexibility is crucial for research that explores complex, evolving questions in investigative toxicology.
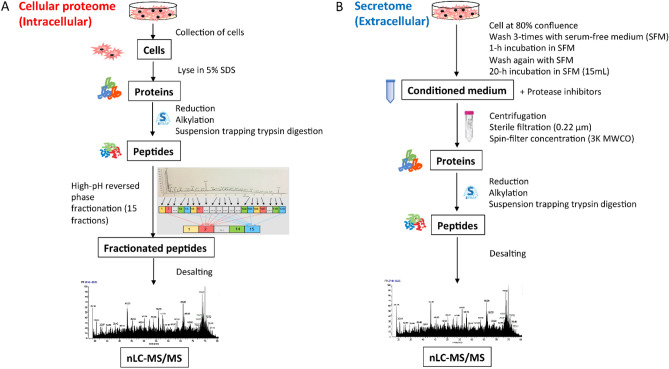
Figure 3. Schematic Workflow for Quantifying Cellular Proteome (A) and Secretome (B) [3]
2. Proteomics' Critical Role in Characterizing Novel Protein Degradation Agents
Mass spectrometry-based proteomics analysis provides a discovery tool for understanding candidate drugs' mechanisms of action. When applied to PROTACs, these techniques offer unbiased insights into binding, degradation selectivity, and mechanisms related to efficacy and safety. Global analyses can identify direct degradation events and assess downstream pathway regulation potentially caused by degradation or off-target inhibition. Targeted proteomics techniques can be used to quantify the levels of relevant E3 ligases and the protein of interest in cell lines and tissues of interest, which can inform the line of sight and provide insights on possible safety liabilities early in the project. Proteomics techniques also help assess protein turnover and resynthesis rates, clarifying target druggability and PK/PD relationships. One study reviewed literature on mass spectrometry-based proteomics in PROTAC development and proposed a proteomics cascade to support targeted protein degradation projects.
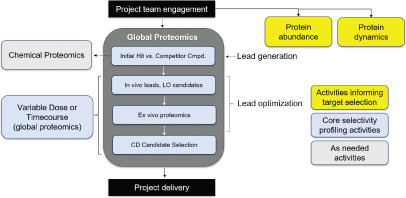
Figure 4. Proteomics Applications for PROTAC Project Prospects [4]
References
[1] Sincere NI, Anand K, Ashique S, Yang J, You C. PROTACs: Emerging Targeted Protein Degradation Approaches for Advanced Druggable Strategies. Molecules. 2023 May 10;28(10):4014. doi: 10.3390/molecules28104014. PMID: 37241755; PMCID: PMC10224132.
[2] Hemkens M, Stamp K, Loberg LI, Moreau K, Hart T. Industry perspective on the nonclinical safety assessment of heterobifunctional degraders. Drug Discov Today. 2023 Aug;28(8):103643. doi: 10.1016/j.drudis.2023.103643. Epub 2023 May 26. PMID: 37244567.
[3] Liu X, Zhang Y, Ward LD, Yan Q, Bohnuud T, Hernandez R, Lao S, Yuan J, Fan F. A proteomic platform to identify off-target proteins associated with therapeutic modalities that induce protein degradation or gene silencing. Sci Rep. 2021 Aug 4;11(1):15856. doi: 10.1038/s41598-021-95354-3. PMID: 34349202; PMCID: PMC8338952.
[4] Zhang AX, Cassidy K, Dahl G, Moreau K, Pachl F, Zuhl AM. The Vital Role of Proteomics in Characterizing Novel Protein Degraders. SLAS Discov. 2021 Apr;26(4):518-523. doi: 10.1177/2472555220985776. Epub 2021 Feb 20. PMID: 33615886.
MtoZ Biolabs, an integrated chromatography and mass spectrometry (MS) services provider.
Related Services
How to order?







