Signal Peptide Prediction Service
- Signal peptide prediction result table (Excel)
- Visualization images of cleavage sites and scoring curves (PNG or PDF)
- Multi-model prediction comparison files (outputs from Neural Network, HMM, SVM, etc.)
- Comprehensive analysis report (PDF), including method descriptions, result interpretation, and recommendations
MtoZ Biolabs has launched the signal peptide prediction service which enables accurate prediction and characterization of signal peptide regions within protein sequences. This service is widely used in recombinant protein expression design, secretion pathway studies, and protein processing mechanism evaluation, providing reliable data support for sequence optimization, structural verification, and functional research.
What Is Signal Peptide?
A signal peptide is a short N-terminal peptide composed of hydrophobic amino acids that directs nascent proteins into the secretory pathway or specific cellular organelles, ensuring proper processing. The presence, length, and cleavage site of a signal peptide can significantly influence protein secretion efficiency, folding stability, and biological function. By predicting signal peptides, researchers can assess secretion potential and localization characteristics in advance, avoid suboptimal expression design, and improve the success rate of protein engineering and functional studies. Therefore, signal peptide prediction plays an important role in recombinant protein production, structural research, and cell biology.
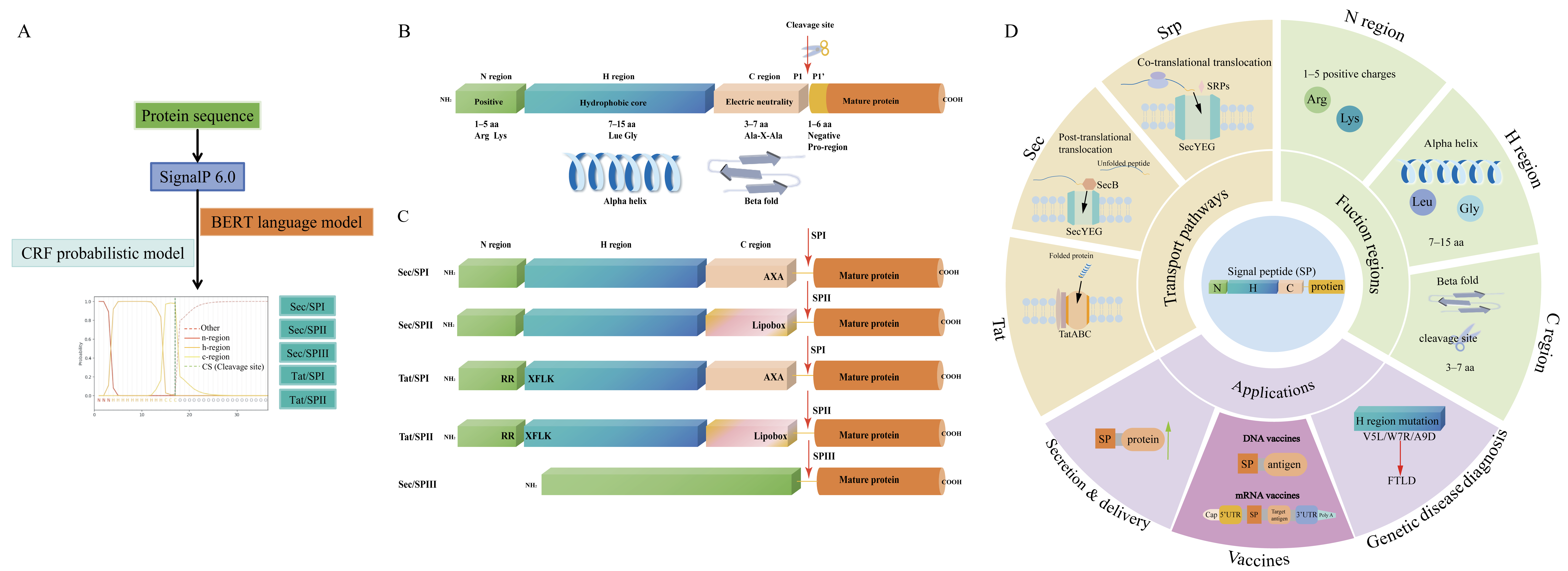
Zhang, S.et al. Biomolecules, 2025.
Figure 1. A Comprehensive Analysis of Signal Peptide Prediction, Structure, and Application.
Signal Peptide Prediction Service at MtoZ Biolabs
1. Neural Network-Based Signal Peptide Prediction
MtoZ Biolabs applies deep neural network models to learn sequence features, enabling high-accuracy identification of signal peptide regions and cleavage sites, suitable for sequence analysis across multiple biological species.
2. Sequence Determination Based on Hidden Markov Models (HMM)
Through HMM statistical modeling of amino acid distribution features, this method reliably predicts signal peptide domains and transport patterns, supporting cross-species conservation studies.
3. Rapid Screening Based on Rule Matrices
Using classical rule matrices to score potential signal sequences enables fast screening of large sets of candidate proteins.
4. Cross-Validation Using Multiple Algorithms
By integrating and comparing results from multiple prediction tools, this approach improves the accuracy of signal peptide identification and cleavage site localization.
5. Classification of Special Types of Signal Peptides
This service supports the analysis of different types of signal peptides, such as Sec, Tat, and Lipoprotein Signal Peptide, making it suitable for studies of secreted proteins, membrane proteins, and specific processing pathways.
Workflow of Signal Peptide Prediction Service
1. Sequence Collection and Quality Check
The protein or nucleotide sequences provided by the client are formatted and examined for completeness to ensure they can be used for signal peptide prediction analysis.
2. Multi-Model Prediction Processing
The sequences are analyzed using multiple models to perform parallel predictions with various algorithms.
3. Identification of Signal Peptide Regions and Cleavage Sites
Outputs from different algorithms are integrated to locate potential signal peptide regions and to evaluate the presence and confidence of cleavage sites.
4. Result Integration and Model Comparison
Prediction results from different methods are cross-validated, and highly consistent information is selected to generate comprehensive scoring and reliability assessment.
5. Report Generation and Functional Interpretation
Based on the final prediction results, an explanatory report is generated that includes signal peptide presence assessment, cleavage site analysis, and potential secretion pathway interpretation for downstream research use.
Why Choose MtoZ Biolabs?
✅ Multi-Algorithm Integration: Integrates neural networks, hidden Markov models, and other prediction tools to achieve more stable and reliable results.
✅ High-Confidence Localization: Performs cross-validation of signal peptide regions and cleavage sites to reduce misclassification and omissions.
✅ Data Quality Assurance: Implements rigorous sequence quality checks to ensure accurate prediction inputs.
✅ Full-Process Support: Provides end-to-end service from sequence preparation and prediction analysis to result interpretation.
✅ Customized Analysis: Delivers tailored analytical strategies based on research objectives and sample characteristics.
Applications of Signal Peptide Prediction Service
1. Recombinant Protein Expression Design
Used to predict the presence and strength of secretion signal peptides, guiding the optimization of secretion expression strategies for exogenous proteins in cellular or microbial systems.
2. Cellular Secretion Pathway Studies
Helps researchers analyze protein secretion localization and transport routes, supporting investigations of extracellular regulation and secretion mechanisms.
3. Protein Function Annotation
Used to preliminarily infer the secretion properties of unknown proteins and assist in subcellular localization prediction and functional classification.
4. Transmembrane Protein Maturation and Targeting Studies
Used to analyze the role of signal peptides in transmembrane protein maturation, cleavage, folding, and targeting, providing support for structural biology research.
5. Biologic Quality Control and Consistency Verification
Used to examine whether signal peptides are completely cleaved and to identify quality issues caused by abnormal expression, improper processing, or incomplete secretion.
Deliverables
1. Comprehensive Experimental Details
2. Signal Peptide Prediction Result Table
3. Sequence Feature Visualization Diagrams
4. Integrated Results of Multi-model Prediction
5. Comprehensive Analysis Report
6. Raw Data Files
FAQ
Q1: What types of samples are suitable?
A1: This service is applicable to any sample that can provide amino acid sequence information, including gene sequences, transcript sequences, protein sequences, and recombinant protein design drafts. Physical samples are not required; reliable sequence information is sufficient to perform signal peptide prediction and cleavage site analysis.
Q2: What is the service general workflow?
A2:
Q3: What data formats are provided?
A3: The deliverables include:
If special analytical requirements exist, data formats can be customized according to project specifications.
Q4: How should I prepare my samples?
A4: To ensure accurate prediction, it is recommended to:
1. Provide complete and accurate amino acid sequences or coding sequences (FASTA or text format)
2. Indicate mutation positions and sequence versions if recombinant variants or designed proteins are involved
3. Provide relevant background information to optimize the prediction strategy
4. Perform quality checks on sequencing-derived sequences to avoid mismatches or missing regions
For more information, please refer to Sample Submission Guidelines for Proteomics, Sample Submission Guidelines for Metabolomics.
Start Your Project with MtoZ Biolabs
Contact us to discuss your experimental design or request a quote. Whether you are exploring the secretion features of a target protein or evaluating the impact of signal peptides on expression efficiency, MtoZ Biolabs can provide accurate and reliable prediction support.
How to order?







