Protein-peptide Interaction Prediction Service
-
Peptide-based drug discovery and inhibitor design.
-
Structural elucidation of signaling or immune receptor binding.
-
Vaccine epitope identification and validation.
-
Analysis of disease-associated peptide-protein interfaces.
-
Development of peptide scaffolds for molecular targeting.
MtoZ Biolabs provides comprehensive Protein-Peptide Interaction Prediction Services designed to support drug discovery, peptide therapeutics design, and structural biology research. Using advanced computational modeling and structure-based algorithms, we predict and characterize the binding affinities, interaction sites, and molecular mechanisms underlying protein-peptide recognition. This service integrates multiple bioinformatics and simulation platforms, enabling researchers to prioritize candidates with high binding specificity and stability before experimental validation.
What Is Protein-peptide Interaction Prediction?
Protein-peptide interactions are central to regulating cellular signaling, enzyme function, and immune response. Experimentally identifying these interactions often relies on techniques such as co-immunoprecipitation, pull-down assays, or surface plasmon resonance, which are time-consuming, costly, and limited by experimental conditions. In contrast, computational prediction methods provide a faster and more efficient approach for screening large datasets, reducing experimental workload, and guiding downstream validation.
Through computational modeling, it is possible to predict binding regions, assess interaction strength, and evaluate molecular stability before laboratory testing. These prediction-based strategies accelerate early-stage discovery of peptide therapeutics, enzyme modulators, and receptor-binding candidates, significantly enhancing research efficiency and reproducibility.
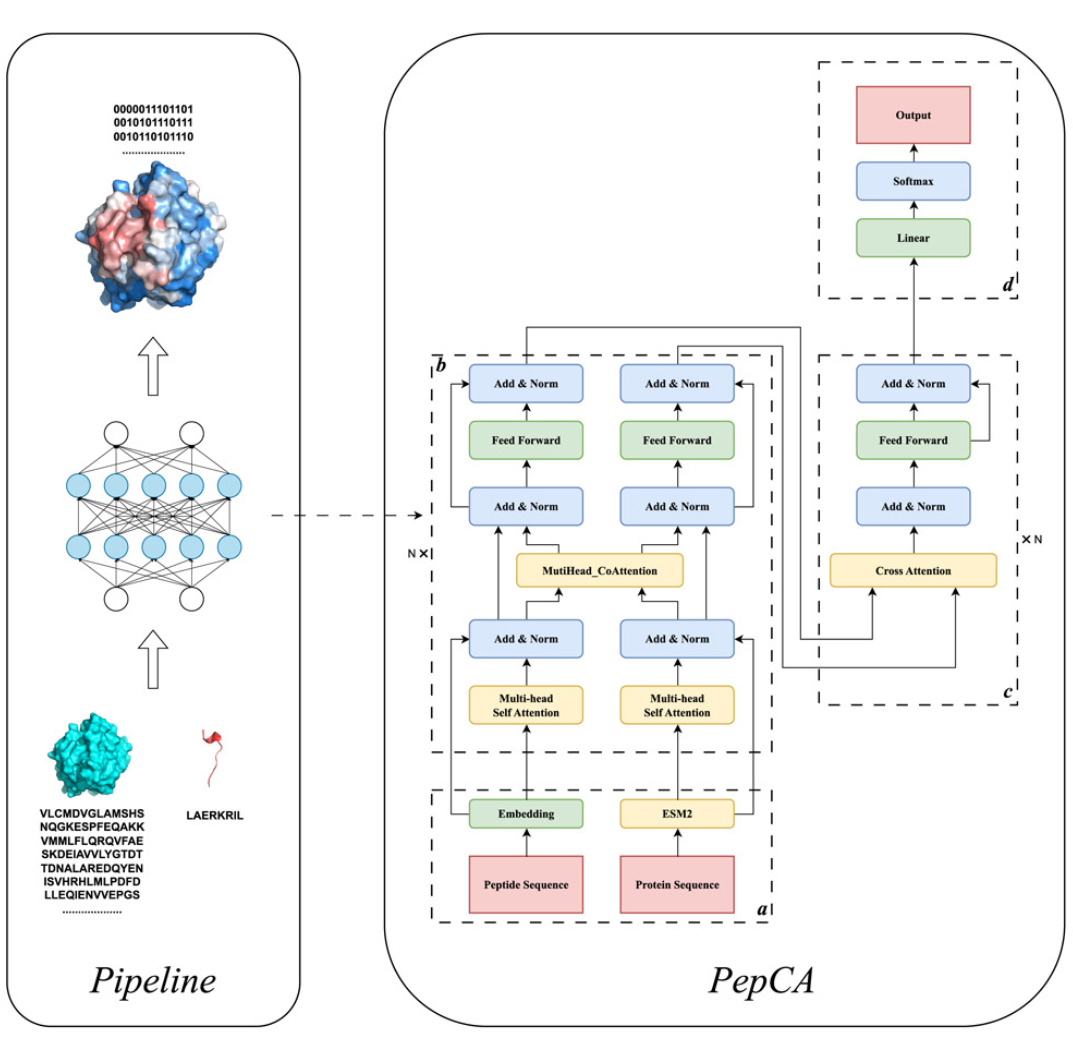
Huang, J. et al. iScience. 2024.
Figure 1. Unveiling Protein-peptide Interaction Sites with a Multi-Input Neural Network Model
Principle of Protein-peptide Interaction Prediction
MtoZ Biolabs integrates multiple computational frameworks to predict and characterize protein-peptide interactions with high accuracy and biological relevance. Our platform combines:
1. Machine Learning Models: Utilizes trained neural networks and quantitative structure-activity relationship (QSAR) features to infer interaction patterns from known datasets.
2. Template-Based Prediction: Identifies conserved motifs and homologous binding templates from protein-peptide interaction databases to guide structural modeling.
3. Nearest Neighbor Algorithm: Compares new protein-peptide pairs with known complexes to predict binding likelihood and interface geometry based on sequence and structure similarity.
Protein-peptide Interaction Prediction Service at MtoZ Biolabs
💠Protein-Peptide Binding Site Identification
Prediction of potential interaction hotspots on protein surfaces and mapping of key residues involved in peptide recognition.
💠Affinity Prediction and Energy Calculation
Quantitative evaluation of binding strength using energy minimization, scoring functions, and thermodynamic free energy calculations.
💠Interaction Mode and Interface Characterization
Visualization and analysis of binding geometry, hydrogen-bond networks, hydrophobic contacts, and salt bridges that stabilize the complex.
💠Dynamic Simulation and Stability Assessment
Time-dependent simulation of protein-peptide complexes to examine flexibility, solvent exposure, and conformational dynamics.
💠Protein-Peptide Network Prediction
Systems-level modeling of peptide-mediated protein interaction networks to identify central regulatory pathways and potential therapeutic targets.
💠Integration with Experimental Design
We provide computational results that guide peptide synthesis, structural validation, and affinity measurements in subsequent experimental stages.
Why Choose MtoZ Biolabs
✔️Proven Expertise
Extensive experience in peptide-protein modeling, computational biology, and structural bioinformatics.
✔️Advanced Computational Platform
Integrates state-of-the-art algorithms and high-performance computing for precise and efficient predictions.
✔️Comprehensive Workflow
Covers every stage from data preprocessing and prediction to visualization and biological interpretation.
✔️Tailored Project Design
Flexible service customized to client goals, sample type, and desired analytical depth.
✔️Dedicated Technical Support
Expert scientists provide one-on-one consultation, ensuring smooth communication and project success.
Applications of Protein-peptide Interaction Prediction Service
FAQ
Q1: What is the service general workflow?
Q2: What data formats are provided?
We provide comprehensive data outputs in multiple formats to ensure easy access and downstream analysis. Standard deliverables include:
1. Detailed result reports in PDF format summarizing workflow, prediction models, and analysis outcomes.
2. Raw and processed data files in CSV or Excel formats for further statistical or computational processing.
3. Structural data files in PDB format and interaction network data in TXT or JSON formats for visualization and modeling.
4. High-resolution graphical outputs in PNG or TIFF for publication or presentation use.
5. Additional formats can be provided upon request to meet specific research or submission requirements.
Start Your Project with MtoZ Biolabs
MtoZ Biolabs combines computational precision with biological insight to deliver reliable, actionable predictions of protein-peptide interactions. Our advanced algorithms, expert team, and customized workflow empower researchers to accelerate discovery, reduce experimental workload, and gain deeper mechanistic understanding.
For more information or to request a customized project consultation, please Contact us today.
How to order?







