Protein-Peptide Binding Regions Prediction Service
MtoZ Biolabs provides Protein-Peptide Binding Regions Prediction Service to help researchers identify, characterize, and prioritize the interaction interfaces between peptides and target proteins using advanced computational modeling. By integrating machine learning algorithms, structural bioinformatics, and fragment based docking simulations, our service offers accurate and data driven predictions that guide peptide design, mechanism studies, and early stage therapeutic evaluation.
Background
Peptides interact with proteins through specific binding regions that involve defined residues, structural motifs, and complementary physicochemical features. Understanding these interaction sites is essential for rational peptide optimization, function prediction, and the development of peptide based therapeutics. Experimental characterization of protein peptide binding regions often requires resource intensive approaches such as mutagenesis, NMR spectroscopy, or crystallography. These methods can be expensive, time consuming, and technically challenging when screening multiple peptide candidates.
Computational prediction offers a powerful and efficient alternative. By leveraging structural data, sequence features, machine learning models, and docking simulations, researchers can rapidly identify potential binding regions without performing large scale experiments. Such predictions provide mechanistic insight into peptide recognition, affinity determinants, and possible competition with endogenous ligands. They also support downstream studies including peptide optimization, binding affinity evaluation, and functional annotation.
Protein-Peptide Binding Regions Prediction Service at MtoZ Biolabs
MtoZ Biolabs delivers a comprehensive computational platform designed specifically for identifying peptide binding regions on protein targets. Using ACCLUSTER clustering algorithms, deep convolutional neural networks, random forest classifiers, and fragment based docking workflows, our service enables high resolution prediction of protein peptide interfaces supported by quantitative scoring metrics and structural visualization.
💠ACCLUSTER-Based Interaction Region Mapping
We use ACCLUSTER clustering algorithms to analyze structural features, classify potential binding pockets, and identify regions with favorable geometric and chemical characteristics. This method improves detection of physically meaningful binding surfaces.
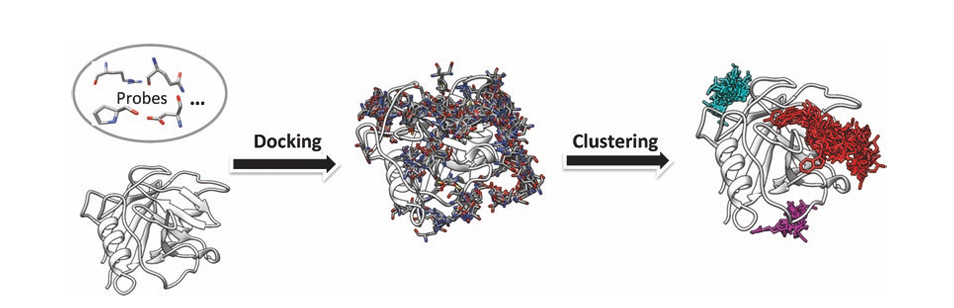
Yan, C. et al. Methods Mol Biol. 2017.
Figure 1. The Schematic Diagram of The ACCLUSTER Method
💠Deep Convolutional Neural Networks (CNN)
Our CNN models are trained on large structural datasets to recognize binding motifs and interaction patterns from protein surfaces. CNN analysis enhances prediction accuracy by capturing complex spatial features that traditional descriptors may overlook.
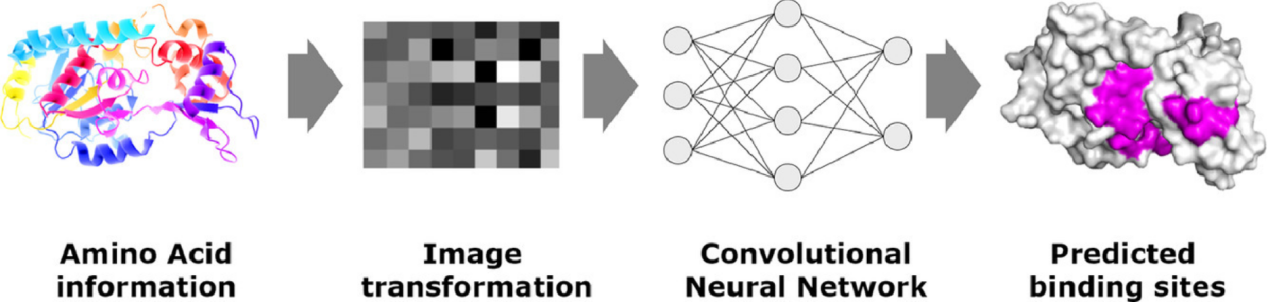
Wardah, W. et al. J Theor Biol. 2020.
Figure 2. Predicting Protein-Peptide Binding Sites with a Deep Convolutional Neural Network
💠Random Forest Classification
Random forest models combine multiple types of features such as surface curvature, residue environment, evolutionary conservation, hydrophobicity, and electrostatic patterns. This multilayer decision process ranks candidate binding regions based on probability scores.
💠Fragment-Based Docking Simulation
We implement fragment based docking simulations to explore how peptide fragments engage with the protein surface. This approach reveals local binding hotspots, evaluates interaction energy, and refines predicted regions using conformational sampling.
💠Integrative Scoring and Consensus Modeling
All modeling results are combined through a consensus scoring framework that improves overall reliability and highlights regions with agreement across multiple prediction methods.
💠Structural Visualization and Annotation
Our team generates clear structural models and annotated protein surfaces displaying predicted binding regions and their associated scores.
Why Choose MtoZ Biolabs
✔️Extensive Experience
Our team has deep expertise in peptide protein interaction analysis and applies proven methodologies to support diverse research objectives.
✔️Reliable Data Outputs
All predictions are supported by transparent scoring metrics, consensus modeling, and clearly annotated structural results.
✔️Customizable Analysis Plans
We tailor every project based on the target protein, peptide characteristics, and study goals, delivering flexible workflows that suit various research stages.
✔️Professional Scientific Team
Our computational biologists provide expert guidance throughout the project and ensure high quality scientific interpretation.
✔️Responsive Client Support
We offer timely communication, dedicated project assistance, and follow up discussions to help clients apply results effectively.
Applications of Protein-Peptide Binding Regions Prediction Service
1. Peptide Optimization
Identify critical binding residues and refine peptide sequences for improved affinity.
2. Screening of Peptide Candidates
Predict binding potential for multiple peptide variants to prioritize candidates.
3. Drug Interaction Analysis
Evaluate whether peptides compete with endogenous ligands or known modulators.
4. Structural Model Development
Generate detailed interaction maps useful for downstream docking or molecular dynamics simulations.
5. Epitope or Motif Discovery
Identify sequence motifs associated with high affinity binding.
6. Mapping Protein Functional Regions
Clarify protein functional domains involved in modulation or signaling.
FAQ
Q1: What is the service general workflow?
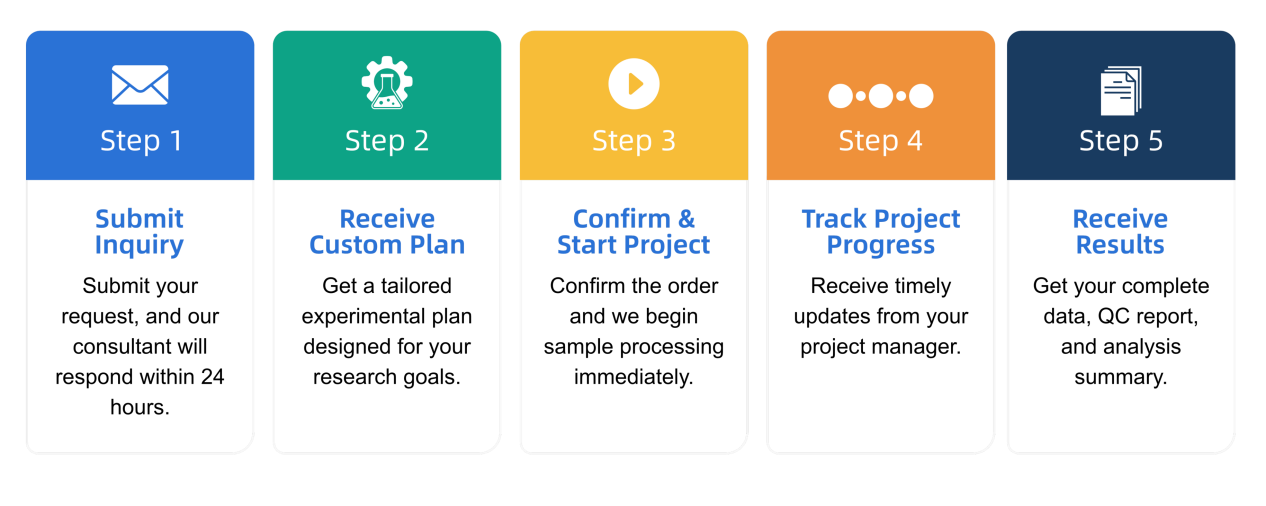
Q2: What data formats are provided?
We supply all prediction results and structural outputs in standard formats suitable for downstream analysis:
1. PDF report summarizing methods, prediction results, confidence scores, and interpretation.
2. PDB or mmCIF files for predicted binding poses, structural models, and docking conformations.
3. Excel or CSV files containing residue level scores, binding probability tables, and ranked binding region lists.
4. High resolution PNG or TIFF images showing structural visualizations, and model highlights.
5. Additional formats can be provided to meet specific publication, modeling, or regulatory needs.
Start Your Project with MtoZ Biolabs
MtoZ Biolabs provides a comprehensive Protein Peptide Binding Regions Prediction Service using advanced computational methods to help researchers understand and optimize peptide protein interactions. Our multi method platform integrates deep learning, feature based classification, clustering algorithms, and fragment based docking to deliver accurate, interpretable predictions.
Contact us to design a customized analysis plan tailored to your peptide development needs.
How to order?







