Peptide Binding Site Prediction Service
-
Nearest Neighbor algorithms for rapid similarity-based inference
-
Support Vector Machines for high-dimensional feature classification
-
Decision Trees for interpretable interaction pattern analysis
-
Bayesian Networks for probabilistic modeling of residue interactions
-
Neural Networks for nonlinear feature extraction and complex pattern learning
-
Ensemble Learning methods to combine multiple model outputs for superior predictive performance
MtoZ Biolabs provides comprehensive Peptide Binding Site Prediction Service to support peptide drug discovery, target validation, structural biology, and computational molecular design. Our service integrates advanced computational modeling, structural bioinformatics, and data-driven prediction strategies to identify peptide binding interfaces on proteins with high specificity and precision.
What Is Peptide Binding Site Prediction?
Peptides function as highly selective and versatile therapeutic molecules. Their interactions with protein targets determine receptor activation, enzyme modulation, structural stabilization, and inhibition mechanisms. Accurately locating the binding interface enables scientists to design more potent peptide ligands, optimize affinity, improve selectivity, and predict downstream biological effects.
Peptide binding site prediction refers to the computational identification of specific regions on a protein surface that are most likely to interact with a peptide ligand. These regions often include surface grooves, pockets, or exposed residues that provide the structural and chemical features necessary for peptide recognition.
Experimental methods such as mutagenesis, NMR, and crystallography remain valuable but often require substantial resources, long development cycles, and complex sample preparation. Computational binding site prediction provides a faster, more flexible, and cost-effective means to screen possible binding regions before experimental validation, thus accelerating discovery workflows and reducing development risk.
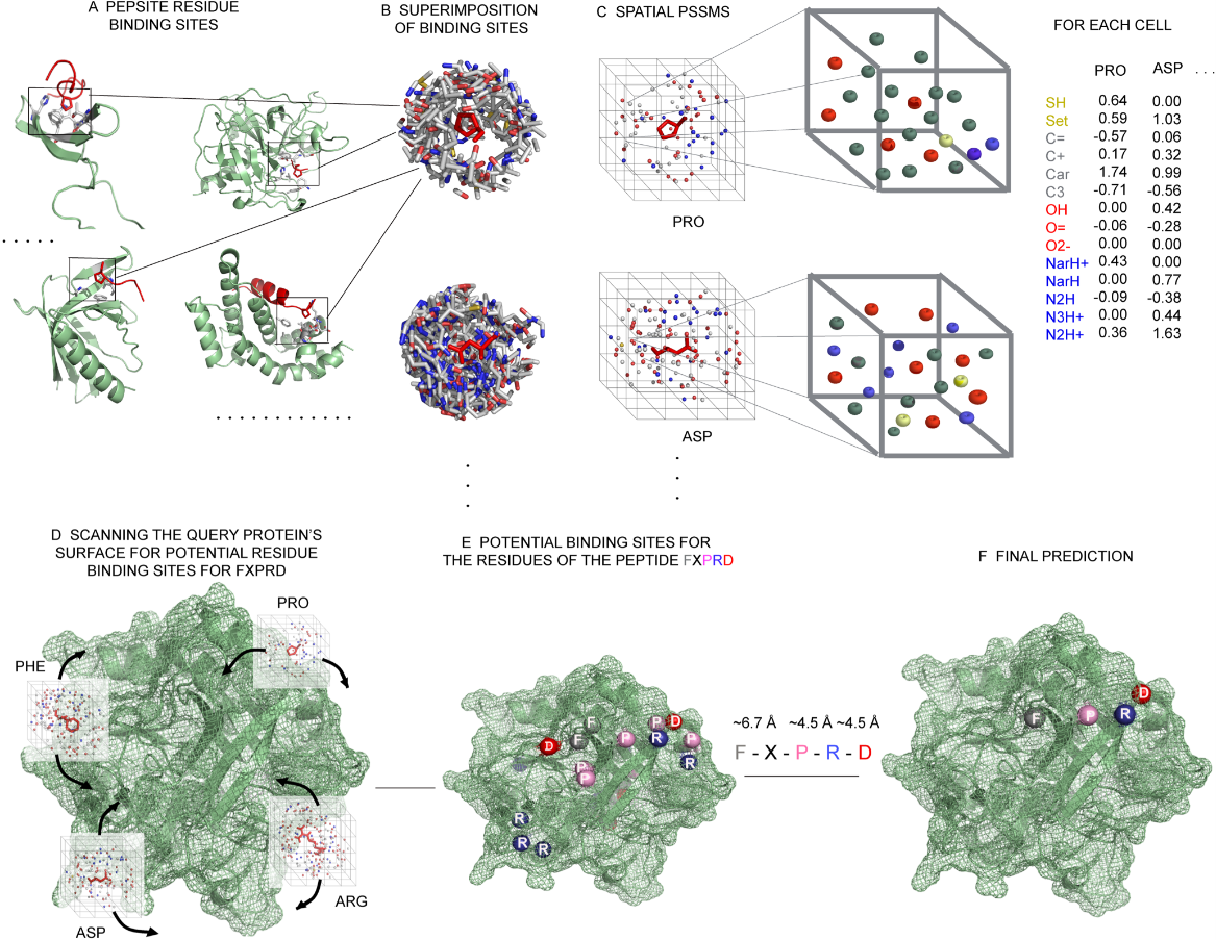
Petsalaki, E. et al. PLoS Comput Biol. 2009.
Figure 1. Overview of the Method of Peptide Binding Site Prediction
Peptide Binding Site Prediction Service at MtoZ Biolabs
Peptide binding site prediction integrates sequence information, structural analysis, docking evaluation, and advanced machine learning to identify protein regions that can interact with peptide ligands.
1. Sequence-Based Prediction
This strategy analyzes primary protein sequences to identify regions that display evolutionary conservation, characteristic residue patterns, surface exposure, physicochemical features, and motif signatures associated with peptide recognition. Comparative sequence analysis and feature extraction help identify candidate binding segments even when no structural information is available.
2. Structure-Based Prediction
When three-dimensional protein structures are accessible, structure-based approaches examine spatial conformation, surface topology, electrostatic distribution, and hydrophobic patches to locate peptide-compatible pockets. Structural modeling and homology-derived models further expand prediction coverage for proteins without experimentally solved structures.
3. Energy-Based Docking and Scoring
Computational docking is applied to evaluate the compatibility between peptides and predicted protein regions. Energy scoring functions assess hydrogen bonding, van der Waals interactions, electrostatic forces, and overall binding stability. Refinement steps help determine the most probable binding poses under near-physiological conditions.
4. Machine Learning Models
To enhance prediction robustness, machine learning algorithms trained on curated protein–peptide interaction datasets provide statistical and pattern-based predictions. MtoZ Biolabs applies model architectures selected according to project needs, including:
Why Choose MtoZ Biolabs
☑️Advanced computational platforms that integrate structural modeling, machine learning and sequence based algorithms for accurate peptide binding site prediction.
☑️Expert team with extensive experience in structure-based modeling, enabling reliable identification of functional surfaces and interaction hotspots.
☑️Flexible workflows tailored to different project goals, including early discovery, binding mechanism studies, and downstream experimental design.
☑️High quality data outputs supported by rigorous internal quality control and transparent reporting.
☑️Responsive scientific support that provides interpretation, follow up consultation and guidance for next step experimental validation.
Applications of Peptide Binding Site Prediction Service
Our Peptide Binding Site Prediction Service supports a broad range of applications across structural biology, molecular design, target validation, and therapeutic peptide discovery.
1. Peptide Drug Discovery
Identification of binding interfaces accelerates rational peptide design.
2. Mechanistic Studies
Binding site information clarifies regulatory mechanisms and protein function.
3. Enzyme Inhibition Research
Peptide inhibitors often target catalytic or allosteric regions identified by prediction.
4. Receptor–Ligand Studies
Prediction helps determine peptide recognition domains for receptors and signaling proteins.
5. Antigen and Epitope Discovery
Mapping binding regions supports immunology and vaccine research.
FAQ
Q1: What is the service general workflow?
Q2: What data formats are provided?
We provide prediction reports in PDF format, structural models in PDB format, and docking or scoring outputs in CSV or TXT format. Visualization files compatible with molecular graphics programs are also included. Additional formats can be provided upon request to meet specific project requirements.
Start Your Project with MtoZ Biolabs
Our peptide binding site prediction framework enhances decision making, reduces experimental burden, and provides a strategic foundation for subsequent validation studies.
If you are interested in our Peptide Binding Site Prediction Service, please feel free to Contact us.
How to order?







