Peptide Array-Based Epitope Mapping Service
- Detection of binding ability for peptide segments covering the target antigen sequence
- Preliminary localization of linear epitopes and screening of key regions
- Comparative analysis of binding characteristics across different antibodies or receptors
- Semi-quantitative evaluation and visualization of binding signals
- Epitope mapping result table (Excel format)
- Array scan images, heatmaps, and other visualized figures (TIFF or PNG format)
- Raw array signal or reading files (depending on the experimental platform)
- Comprehensive analysis report (PDF), including experimental workflow, data interpretation, and key result descriptions
MtoZ Biolabs has launched the peptide array-based epitope mapping service which enables systematic screening and precise localization of potential linear epitope regions within target antigen sequences. This service is widely applied in antibody development, vaccine design, and protein interaction studies, providing reliable data support for epitope identification, binding mechanism analysis, and functional region determination.
Principle of Peptide Array
Peptide array is a high-throughput detection platform that arranges large numbers of short peptides with defined sequences on membranes or chips, enabling rapid screening of antibody or receptor binding regions. Its fundamental principle involves synthesizing peptides that cover the target protein sequence and detecting their binding signals with antibodies to identify linear epitopes within the antigen. This technology offers advantages such as rapid operation, high throughput, and low sample consumption, and is widely used for antibody screening, immunogenicity analysis, and preliminary localization of functional regions, providing important references for subsequent mechanistic studies and molecular design.
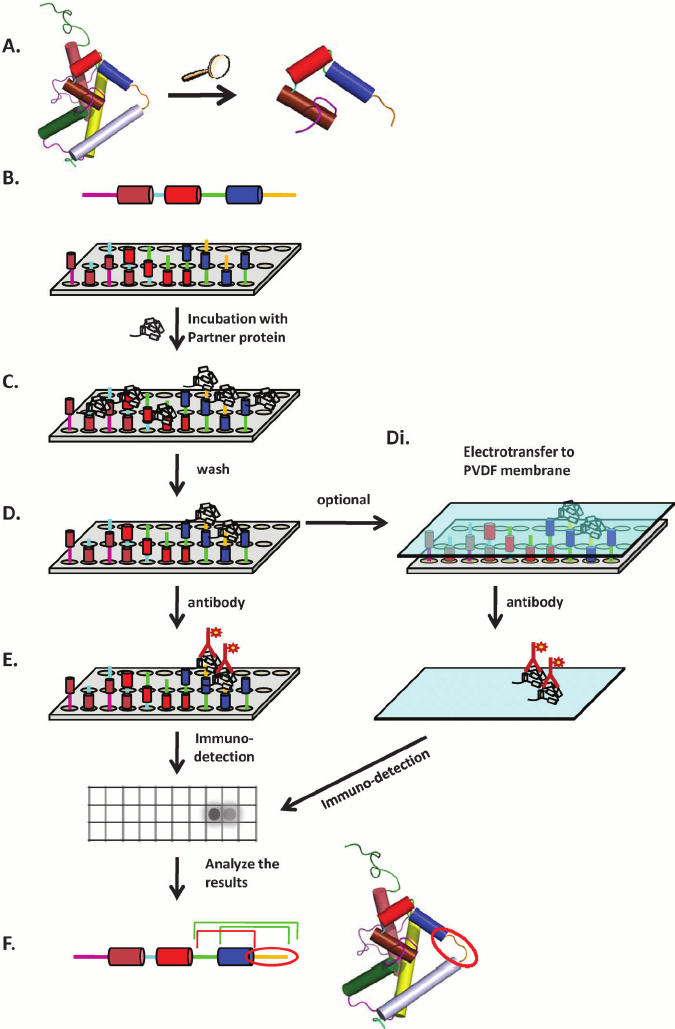
Katz, C. et al. Chemical Society Reviews, 2011.
Figure 1. A General Scheme of Peptide Array Design and Screening.
Peptide Array-Based Epitope Mapping Service at MtoZ Biolabs
MtoZ Biolabs uses high-throughput peptide array (Peptide Array) technology to provide systematic identification and precise localization of linear epitopes within target protein sequences, enabling rapid screening of potential antibody-binding regions.
Our analysis offers comprehensive insights into the immune recognition features of target proteins, helping researchers efficiently define critical epitope regions and supporting downstream mechanistic studies and molecular design.
Workflow of Peptide Array-Based Epitope Mapping Service
1. Peptide Sequence Design
(1) Coverage Design: Generate continuous scanning peptides based on the target protein sequence to ensure complete coverage of potential epitope regions.
(2) Optimized Selection: Use bioinformatics tools to select more representative sequences and improve mapping efficiency.
2. Peptide Array Preparation
(1) Solid-Phase Construction: Fix peptides onto chip or membrane surfaces through solid-phase synthesis or in situ photochemical methods.
(2) Array Layout: Arrange peptides in a high-density spot matrix to enable high-throughput detection.
3. Binding Detection
(1) Antibody Incubation: Incubate samples with the peptide array and identify binding peptides through fluorescence, chemiluminescence, or enzyme-linked detection systems.
(2) Multi-Platform Response Measurement: Combine ELISA, SPR, and other techniques to evaluate binding strength, specificity, and relative affinity.
4. Data Analysis
(1) Signal Interpretation: Identify highly reactive short peptides and locate potential epitope regions.
(2) Result Validation: Compare identified sequences with existing epitope databases to confirm key binding motifs.
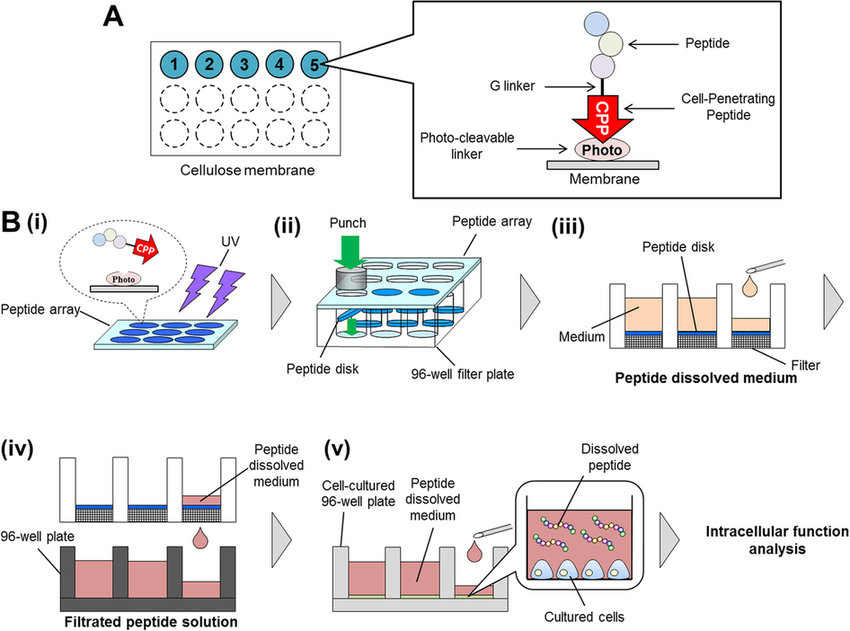
Matsumoto, R. et al. Scientific Reports, 2015.
Figure 2. Schematic Representation of Peptide Array-Based Intracellular Functional Peptide Screening.
Why Choose MtoZ Biolabs?
✅ High-Throughput Screening: Evaluate large numbers of peptides in a single run to improve epitope mapping efficiency.
✅ High-Resolution Analysis: Precisely pinpoint short peptides or core amino acid sequences.
✅ Strong Reproducibility: Standardized array systems ensure stable and reliable signal outputs.
✅ Rapid Analysis Cycle: Efficient array construction and data acquisition accelerate the research process.
✅ Comprehensive One-Stop Support: Covers the full workflow from peptide design and array preparation to detection and data analysis.
Applications of Peptide Array-Based Epitope Mapping Service
1. Antibody Binding Region Identification
Used to rapidly identify linear binding segments between antibodies and antigens, supporting antibody specificity verification and mechanism studies.
2. Protein Structure-Function Analysis
By analyzing epitope distribution, this approach helps assess functional properties and interaction characteristics of different protein regions.
3. Antibody Engineering and Affinity Optimization
Evaluates the binding features of different peptide segments to optimize antibody selectivity and binding performance.
4. Biomarker-Related Peptide Verification
Used to screen antigenic peptide fragments with potential diagnostic value, supporting biomarker target research.
5. Peptide Drug Target Segment Identification
Locates potential functional peptide regions, providing experimental evidence for peptide drug development, optimization, and mechanism research.
Deliverables
1. Comprehensive Experimental Details
2. Materials, Instruments, and Methods
3. Epitope Mapping Result Table
4. Visualization Images Generated From Peptide Array Detection
5. Bioinformatics Analysis
6. Raw Data Files
FAQ
Q1: What types of samples are suitable?
A1: This service is suitable for various sample types, including antibodies, purified proteins, recombinant proteins, and cell-derived antigen materials. To ensure accurate epitope mapping results, samples should have high purity, be free of salts, surfactants, and other interfering impurities, and maintain structural stability.
Q2: What is the service general workflow?
A2:
Q3: What data formats are provided?
A3: The deliverables include multiple standardized data formats, including:
If special analytical requirements exist, data formats can be customized according to project specifications.
Q4: How should I prepare my samples?
A4: To ensure experimental quality, sample preparation is recommended to follow these requirements:
(1) Sample Purity: Avoid interference from high salt, strong buffers, detergents, and precipitates
(2) Storage Conditions: Store at -20℃ for short-term use, and -80℃ for long-term preservation
(3) Transportation: Ship using cold-chain or dry ice to maintain sealing and stability
(4) Supplementary Information: Provide sample concentration, source, antibody or antigen type, and research objectives to optimize the array-based epitope screening strategy
For more information, please refer to Sample Submission Guidelines for Proteomics, Sample Submission Guidelines for Metabolomics.
Start Your Project with MtoZ Biolabs
Contact us to discuss your experimental design or request a quote. Whether you are exploring key epitope regions of a target antigen or studying antibody binding characteristics, MtoZ Biolabs can provide you with precise and reliable epitope mapping support.
How to order?







