Edman Degradation-Based Peptide Sequencing Service
-
Stepwise sequencing of peptide N-terminal amino acids
-
Verification of synthetic peptide sequences and purity confirmation
-
N-terminal identification of proteolytic fragments
-
Detection of N-terminal blockage, modification, or truncation
-
Structural consistency evaluation and batch-to-batch comparison
-
N-terminal sequencing results (text or Excel format)
-
Sequence analysis report (PDF), including experimental procedures and result interpretation
-
Raw detection data files (provided depending on the instrument platform)
-
Visualization images such as chromatograms or signal intensity curves (TIFF or PNG format)
MtoZ Biolabs has launched the Edman degradation-based peptide sequencing service which enables stepwise analysis and precise determination of the N-terminal amino acid sequence of peptides. This service is suitable for purified peptides, recombinant protein fragments, and sequence confirmation in the quality control of biological products, providing reliable data support for structural verification, functional studies, and consistency assessment in production.
What Is Edman Degradation?
Edman degradation is a classical and highly specific N-terminal peptide sequencing method that uses phenyl isothiocyanate (PITC) to selectively label and sequentially cleave the N-terminal amino acid, converting it into an identifiable PTH-amino acid to read the peptide sequence step by step. This method does not rely on databases and can directly and accurately confirm N-terminal amino acid composition. It is applicable to verifying synthetic peptide sequences, confirming mutation sites, evaluating protein processing completeness, and assessing the consistency of biopharmaceutical products. Edman degradation remains an indispensable technique in peptide sequencing and quality control.
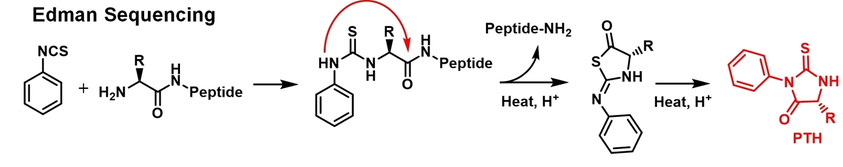
Shi, Q Q. et al. Angewandte Chemie International Edition, 2023.
Figure 1. Edman Degradation for Peptide/Protein Sequencing.
Edman Degradation-Based Peptide Sequencing Service at MtoZ Biolabs
MtoZ Biolabs utilizes Edman degradation sequencing technology to provide high-precision N-terminal peptide sequence analysis, enabling accurate sequence confirmation and assessment of sample quality.
Our sequencing analysis delivers clear and reliable structural information on peptide sequence characteristics, helping researchers accurately assess sample quality and supporting subsequent functional studies and characterization.
Workflow of Edman Degradation-Based Peptide Sequencing Service
1. Sample Evaluation
Assess the purity, N-terminal accessibility, and suitability of the peptide sample to ensure it meets the requirements for Edman degradation analysis.
2. Stepwise N-terminal Degradation
Perform cyclic N-terminal cleavage on the sample, chemically releasing one amino acid derivative per cycle.
3. Derivatized Product Detection
Use a high-resolution separation system to detect each released PTH-amino acid and determine its corresponding amino acid identity.
4. Sequence Reconstruction and Verification
Reconstruct the peptide’s N-terminal sequence based on multi-cycle detection results and confirm sequence continuity and reliability.
Why Choose MtoZ Biolabs?
✅ High accuracy: Stepwise analysis of N-terminal amino acids to obtain true, direct sequence information.
✅ High specificity: No need to rely on fragment ion inference, avoiding ambiguous matches in mass spectrometry sequencing.
✅ Standardized workflow: Excellent reproducibility and high stability to ensure reliable and consistent sequence reading.
✅ Support from a professional technical team: An experienced analysis team ensures accurate and precise interpretation of results.
✅ One-stop service: A one-stop service process covering everything from sample evaluation to final report generation.
Applications of Edman Degradation-Based Peptide Sequencing Service
1. Protein Degradation and Stability Studies
Used to assess protein stability and cleavage patterns under different conditions by identifying the N-terminal sequences of degradation products.
2. Unknown Peptide Structure Identification
Applied to identify unknown peptides, degradation fragments, or impurity components in purified samples, supporting purification process optimization and component confirmation.
3. Biomarker Peptide Analysis
Used to identify specific peptides derived from cell, plasma, or tissue samples to support biomarker exploration and mechanism studies.
4. Natural Product and Bioactive Peptide Identification
Suitable for confirming the sequences of active peptides in extracts from plants, microorganisms, or animal tissues.
5. Antibody/Antigen Characterization and Biopharmaceutical Development
Used to confirm the sequence integrity of recombinant antigens, peptide vaccine components, or antibody fragments, supporting biopharmaceutical quality studies.
Deliverables
1. Comprehensive Experimental Details
2. Materials, Instruments, and Methods
3. N-terminal Sequencing Results
4. Visualization Figures (Such as Chromatograms and Signal Intensity Curves)
5. Sequence Analysis Report
6. Raw Data Files
FAQ
Q1: What types of samples are suitable?
A1: This service is suitable for high-purity peptide or protein samples, especially those requiring N-terminal sequence determination. It is recommended that samples be free of strong buffers, salts, surfactants, and other chemical modifiers, and that they maintain good stability and integrity to ensure sequencing accuracy.
Q2: What is the service general workflow?
A2:
Q3: What data formats are provided?
A3: The deliverables include:
If special analytical requirements exist, data formats can be customized according to project specifications.
Q4: How should I prepare my samples?
A4: To ensure high-quality sequencing results, the following guidelines are recommended:
(1) Sample purity: avoid high-salt conditions, strong buffer systems, detergents, and precipitates
(2) Sample condition: ensure the sample is intact, stable, and free of modifications that may affect N-terminal sequencing
(3) Storage: store at -20℃ for short-term use and at −80℃ for long-term preservation
(4) Shipping: use cold-chain or dry-ice transportation and avoid repeated
freeze-thaw cycles
(5) Supporting information: provide sample concentration, source, and sequencing requirements to design an optimal sequencing plan
For more information, please refer to Sample Submission Guidelines for Proteomics, Sample Submission Guidelines for Metabolomics.
Start Your Project with MtoZ Biolabs
Contact us to discuss your experimental design or request a quote. Whether you aim to verify sequence accuracy or conduct more in-depth peptide structural research, MtoZ Biolabs can provide precise and reliable technical support.
How to order?







