Targeted Ubiquitination Site Analysis Service
MtoZ Biolabs provides Targeted Ubiquitination Site Analysis Service that enables precise identification of ubiquitination sites on specific proteins of interest. This service is designed for researchers aiming to understand how ubiquitin modification affects the activity, stability, and interactions of selected proteins. By integrating selective peptide enrichment with high-resolution LC-MS/MS, MtoZ Biolabs offers accurate, site-level characterization of ubiquitination that supports both mechanistic studies and biopharmaceutical research.
Overview
Ubiquitination is a post-translational modification in which the small regulatory protein ubiquitin is covalently attached to lysine residues of substrate proteins. It acts as a versatile cellular signal that regulates diverse biological processes, including protein degradation, signal transduction, DNA repair, and receptor internalization. The biological consequence of ubiquitination depends on both the modified protein and the type of ubiquitin linkage involved.
Different classes of proteins undergo ubiquitination to achieve distinct regulatory outcomes:
Cell cycle regulators and degradation substrates: Proteins such as cyclins, p53, and p27 are modified through K48-linked polyubiquitination, marking them for proteasomal degradation and ensuring cell cycle control.
Signaling and immune pathway proteins: Adaptor proteins and kinases such as TRAF6 and RIP1 are modified via K63-linked ubiquitination, promoting NF-κB activation and innate immune signaling, while K48-linked ubiquitination of IκBα triggers its degradation to release NF-κB.
Chromatin and DNA repair proteins: Histones H2A and H2B, along with DNA repair proteins like PCNA and FANCD2, are mono- or multi-ubiquitinated to modulate chromatin accessibility and coordinate DNA damage response.
Membrane receptors and trafficking proteins: Receptors including EGFR and Notch are mono-ubiquitinated to regulate endocytosis and lysosomal sorting, thereby modulating signal duration and receptor recycling.
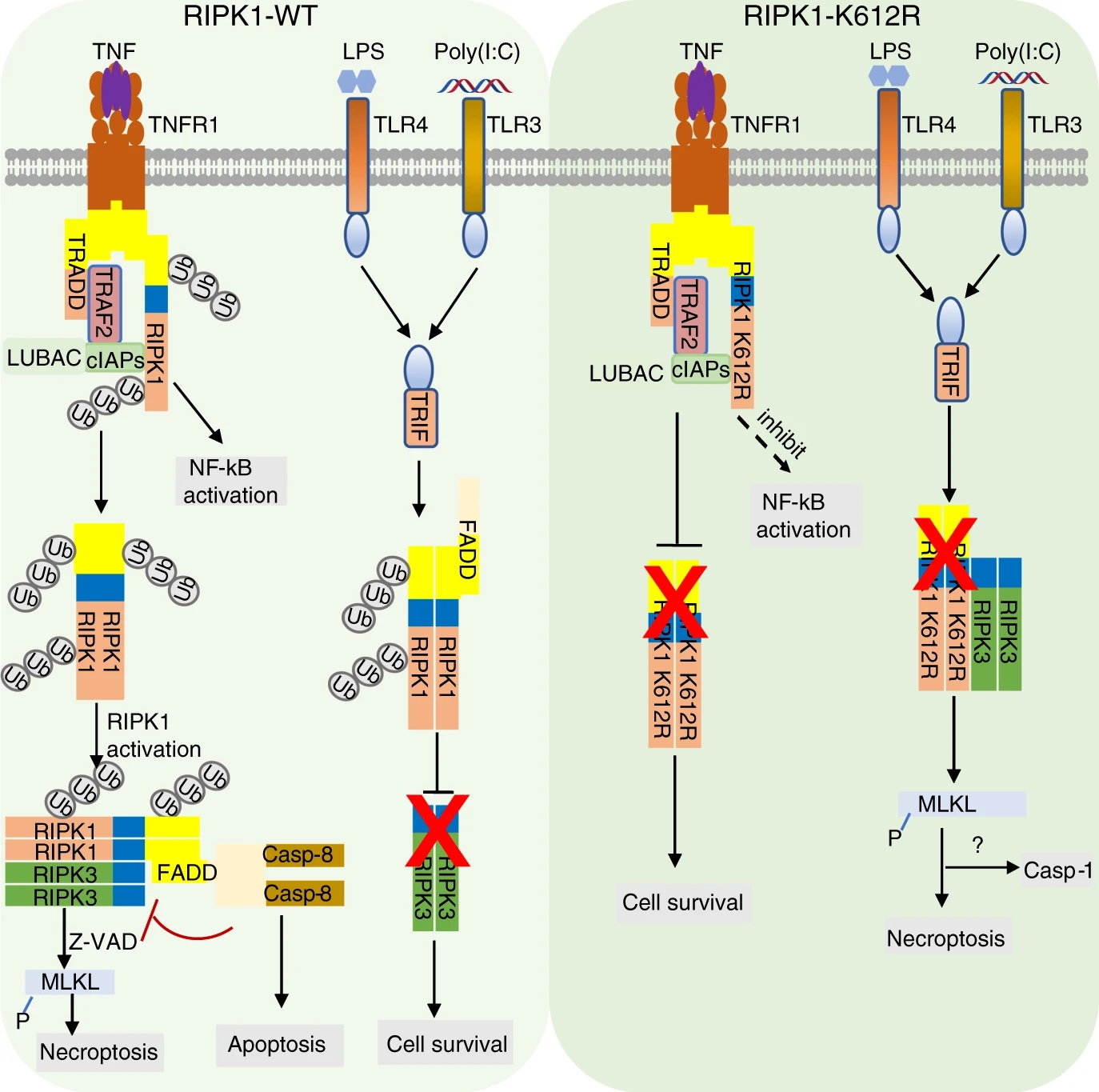
Li, X. et al. Nat. Commun. 2020.
Figure 1. A Model for RIPK1 K612 Ubiquitination in TNFR1 and TLR3/4 Signalings
Targeted Ubiquitination Site Analysis Service at MtoZ Biolabs
MtoZ Biolabs provides an end-to-end analytical solution for mapping and characterizing ubiquitination sites. Each project is customized to the client’s target protein, ensuring accurate and reproducible identification of modification sites.
Our service includes:
Site-Specific Ubiquitination Mapping
Identification of modified lysine residues at amino acid resolution using high-resolution tandem mass spectrometry.
Ubiquitin Linkage Type Characterization
Differentiation of linkage types (e.g., K48, K63, K11) to determine degradation or signaling relevance.
Ubiquitination Site Occupancy Assessment
Evaluation of modification stoichiometry to estimate the degree of ubiquitin attachment at specific sites.
Functional Site Annotation
Integration of identified ubiquitination sites with known protein domains, signaling pathways, and biological processes.
Workflow of Targeted Ubiquitination Site Analysis Service
MtoZ Biolabs applies a systematic and high-sensitivity workflow optimized for targeted ubiquitination site detection.
1. Protein Preparation and Digestion
Samples containing the target protein are reduced, alkylated, and digested into peptides under denaturing conditions to preserve ubiquitin remnants.
2. Ubiquitinated Peptide Enrichment
Enrichment is performed using anti-diGly (K-ε-GG) antibody-based immunoaffinity capture, which specifically recognizes the di-glycine remnants left on lysine residues after trypsin digestion, ensuring selective isolation of ubiquitinated peptides.
3. LC-MS/MS Analysis
Enriched peptides are analyzed using Orbitrap Fusion Lumos or Q Exactive HF systems for precise detection and site localization.
4. Data Analysis and Validation
Spectral data are processed with advanced algorithms to confirm site identity, linkage type, and localization confidence.
5. Comprehensive Reporting
Results are delivered with annotated site tables, structural mapping, and biological interpretation to support functional studies.
Why Choose MtoZ Biolabs?
✔ High-Resolution Site Mapping: Advanced LC-MS/MS instruments provide confident identification and accurate localization of ubiquitination sites.
✔ Optimized Antibody Enrichment: Anti-diGly antibody capture ensures selective isolation of ubiquitinated peptides with minimal background interference.
✔ Comprehensive Data Interpretation: Each dataset is analyzed with domain mapping and pathway enrichment to reveal the biological significance of modification sites.
✔ Customized Experimental Design: Every project is tailored to the user’s specific proteins and research objectives.
✔ Expert Technical Support: Our experienced proteomics specialists provide professional guidance from experimental setup to final data interpretation.
✔ One-Time-Charge: Our pricing is transparent, no hidden fees or additional costs.
Sample Submission Guidelines
To ensure optimal data quality, please prepare samples following these guidelines:
|
Sample Type |
Recommended Amount |
Storage Conditions |
Notes |
|
Purified protein |
≥50 µg |
−80°C |
Provide purity and buffer composition |
|
Cell lysate |
≥500 µg total protein |
Avoid detergents; freeze immediately |
|
|
Tissue extract |
≥50 mg |
Clarify before freezing |
|
|
Recombinant protein |
≥50 µg |
Provide expression system and tag details |
For more information, please refer to Sample Submission Guidelines for Proteomics.
Applications of Targeted Ubiquitination Site Analysis Service
✅ Studying protein degradation mechanisms and proteasome-mediated regulation.
✅ Investigating signal transduction pathways modulated by non-degradative ubiquitination.
✅ Exploring DNA repair and chromatin remodeling mechanisms.
✅ Understanding immune signaling and inflammation regulation.
✅ Supporting drug discovery and PROTAC development through site-specific modification insights.
Start Your Project with MtoZ Biolabs
MtoZ Biolabs provides comprehensive solutions for site-specific ubiquitination analysis, enabling researchers to uncover the molecular mechanisms regulating their target proteins. Our integrated platform, combining precise enrichment and high-resolution mass spectrometry, ensures accurate and reproducible site-level information.
Contact us today to discuss your experimental design or request a customized quotation. Our experts will help you achieve confident ubiquitination site identification tailored to your research goals.
What Could be Included in the Report?
1. Comprehensive Experimental Details
2. Materials, Instruments, and Methods
3. Total Ion Chromatogram & Quality Control Assessment
4. Data Analysis, Preprocessing, and Estimation
5. Bioinformatics Analysis
6. Raw Data Files
Related Services
How to order?







