Targeted Ubiquitination Analysis Service
MtoZ Biolabs provides Targeted Ubiquitination Analysis Service to accurately identify, quantify, and structurally characterize ubiquitination on targeted proteins. This service enables researchers to uncover how ubiquitin modifications regulate protein degradation, signaling, and functional stability. Through advanced LC-MS/MS platforms and anti-diGly antibody-based enrichment workflows, MtoZ Biolabs offers comprehensive and reproducible ubiquitination profiling for mechanistic and translational research.
Overview
Ubiquitination is a key post-translational modification in which the small protein ubiquitin is covalently attached to lysine residues of target proteins by a cascade of E1 activating, E2 conjugating, and E3 ligating enzymes. This modification can determine protein degradation through the proteasome, alter subcellular localization, or influence interaction networks and signaling cascades.
Studying ubiquitination on specific proteins reveals how this modification regulates biological function and disease processes. For example, p53 ubiquitination modulates tumor suppression and protein turnover, while EGFR and NF-κB ubiquitination control signal transduction and inflammatory responses. MtoZ Biolabs combines optimized enrichment strategies and high-resolution LC-MS/MS to deliver site-specific and quantitative insights into ubiquitination events that drive cellular regulation.
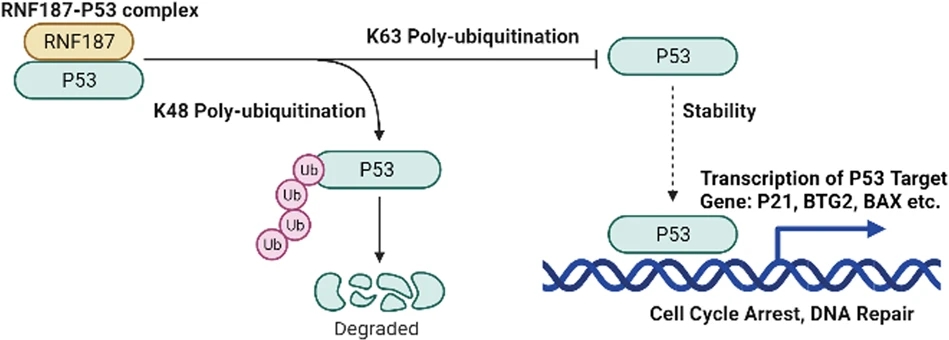
Li, X. et al. CELL DEATH DIS. 2022.
Figure 1. Dual Roles of RNF187-Mediated Polyubiquitination in Regulating p53 Stability and Function
Targeted Ubiquitination Analysis Service at MtoZ Biolabs
MtoZ Biolabs provides a fully integrated workflow for targeted ubiquitination analysis, including peptide enrichment, site identification, quantitative profiling, and structural characterization. Each analytical phase is optimized to ensure high specificity, sensitivity, and reproducibility for confident site-level mapping.
1. Ubiquitinated Peptide Enrichment
Ubiquitinated peptides are selectively enriched using anti-diGly (K-ε-GG) antibody-based immunoaffinity capture, which ensures specific isolation of lysine residues modified by ubiquitin. This enrichment significantly enhances detection sensitivity, enabling comprehensive coverage of low-abundance ubiquitination sites in complex proteomes.
2. Targeted Ubiquitination Identification
High-resolution LC-MS/MS analysis allows precise identification and localization of ubiquitination sites. The workflow distinguishes between mono-, multi-, and poly-ubiquitination and determines major chain linkage types such as K6, K11, K27, K48, and K63.
3. Targeted Ubiquitination Quantification
Quantitative profiling measures relative changes in ubiquitination levels under different biological or experimental conditions. Both label-free and isotopic labeling approaches are available to ensure reproducible, statistically validated quantification for comparative studies.
4. Targeted Ubiquitination Structural Characterization
To explore the structural consequences of ubiquitination, LC-MS/MS is combined with top-down MS, crosslinking MS, or HDX-MS. This analysis characterizes ubiquitin chain topology, linkage heterogeneity, and modification-induced conformational changes, revealing how ubiquitination influences protein structure, stability, and complex formation.
Why Choose MtoZ Biolabs?
✔ Advanced LC-MS/MS Systems: Equipped with Thermo Fisher Orbitrap Fusion Lumos and Q Exactive HF platforms for high-sensitivity, high-resolution ubiquitination analysis.
✔ Specific Anti-diGly Enrichment: Optimized immunoaffinity capture ensures selective isolation of ubiquitinated peptides, maximizing detection efficiency.
✔ Comprehensive Analytical Workflow: From enrichment to structural characterization, each step is seamlessly integrated for consistent, reproducible outcomes.
✔ Quantitative and Comparative Capability: Robust label-free and labeling approaches enable accurate differential quantification of ubiquitination across conditions.
✔ Bioinformatics-Driven Interpretation: In-depth data processing includes site localization scoring, quantification, and pathway enrichment for meaningful biological insight.
Applications of Targeted Ubiquitination Analysis Service
Targeted ubiquitination analysis is essential for understanding protein regulation and cellular signaling. It supports:
Protein degradation and stability research
Signal transduction pathway analysis
DNA damage and repair studies
Cancer biology and immune regulation
Therapeutic target validation and biomarker discovery
Start Your Project with MtoZ Biolabs
MtoZ Biolabs delivers high-resolution, quantitative, and structural characterization of protein ubiquitination with precision and reproducibility
Contact us today to discuss your project or request a customized quotation. Our experts will design a targeted workflow that provides high-quality data and actionable insights for your research and development goals.
FAQ
Q1: What types of samples are suitable?
MtoZ Biolabs accepts a broad range of biological and purified samples for targeted ubiquitination analysis, including cell lysates, tissue extracts, recombinant proteins, and synthetic peptides. Both native and post-translationally modified proteins can be analyzed to identify and characterize ubiquitination sites on specific targets. Recommended sample amounts are ≥500 µg total protein for lysates or ≥50 µg purified protein. MtoZ Biolabs can also assist with protein extraction and purification from complex biological samples when required.
Q2: What is the service general workflow?
Q3: How should I prepare my samples?
Samples should be free of interfering substances such as salts, detergents, or nucleic acids to ensure optimal LC-MS/MS performance. For purified proteins, please provide samples in a compatible buffer with minimal additives. For cell or tissue lysates, ensure efficient lysis and debris removal through centrifugation. For detailed preparation and shipping requirements, please refer to Sample Submission Guidelines for Proteomics.
Q4: What data formats are provided?
MtoZ Biolabs provides comprehensive deliverables to support your research, including:
Raw LC-MS/MS data files
Processed identification and site mapping tables (Excel or CSV format)
Annotated spectra and peptide summary reports (PDF format)
Customized quantitative and interpretive reports according to project objectives (PDF format)
Related Services
How to order?







