Targeted SUMOylation Site Analysis Service
MtoZ Biolabs provides Targeted SUMOylation Site Analysis Service that focuses on identifying and quantifying SUMO modification sites on specific proteins provided by clients. This service enables precise characterization of lysine residues modified by SUMO, supporting advanced studies on protein stability, nuclear localization, and cellular stress response. Utilizing high-resolution LC-MS/MS and specialized SUMO-remnant enrichment workflows, MtoZ Biolabs provides accurate and reproducible site-level data for mechanistic research and biopharmaceutical development.
Overview
SUMOylation is a reversible post-translational modification occurring on lysine residues, regulating protein function, subcellular localization, and interaction networks. Site-specific SUMOylation analysis allows researchers to determine how modification at defined residues influences molecular behavior and cellular processes.
For instance, SUMOylation of PCNA (Proliferating Cell Nuclear Antigen) at lysine 127 and 164 plays distinct roles in DNA repair regulation. SUMO modification prevents unwanted recombination and supports replication fork stability, while ubiquitination at the same region promotes template switching and translesion synthesis, highlighting the complexity of site-level modification events.
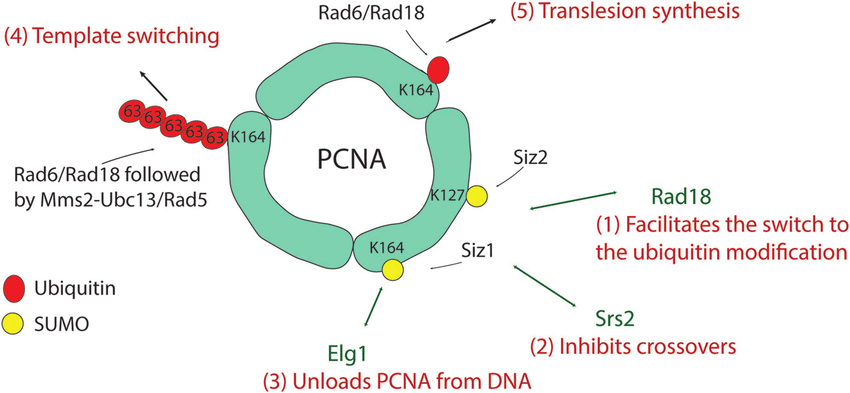
Jalal, D. et al. NAR. 2017.
Figure 1. SUMOylation Sites on PCNA and Their Regulatory Roles in DNA Replication and Repair.
Targeted SUMOylation Site Analysis Service provides a detailed understanding of these specific modification events, delivering residue-level insights essential for protein function characterization and regulatory studies.
Workflow of Targeted SUMOylation Site Analysis
MtoZ Biolabs offers a comprehensive workflow for targeted SUMOylation site analysis, integrating peptide purification, immunoaffinity enrichment, and high-resolution LC-MS/MS to ensure precise and reproducible site identification.
1. Protein Digestion
Protein samples are digested using Lys-C or other suitable proteases under denaturing conditions to generate peptides while preserving SUMO-conjugated residues.
2. SUMO Peptide Purification
Peptides are pre-purified to remove contaminants and subsequently enriched via SUMO-specific immunoprecipitation, ensuring the selective capture of SUMO-modified peptides.
3. Peptide Fractionation
Enriched peptides are fractionated to reduce sample complexity and improve chromatographic resolution, facilitating deeper proteome coverage during LC-MS/MS analysis.
4. LC-MS/MS Analysis
Purified peptide fractions are analyzed using Thermo Fisher Orbitrap Fusion Lumos or Q Exactive HF systems, providing high mass accuracy and detailed site localization of SUMO modifications.
5. Data Processing and Site Mapping
Mass spectrometry data are processed using advanced bioinformatics pipelines to identify SUMO-modified peptides and map modification sites with high confidence.
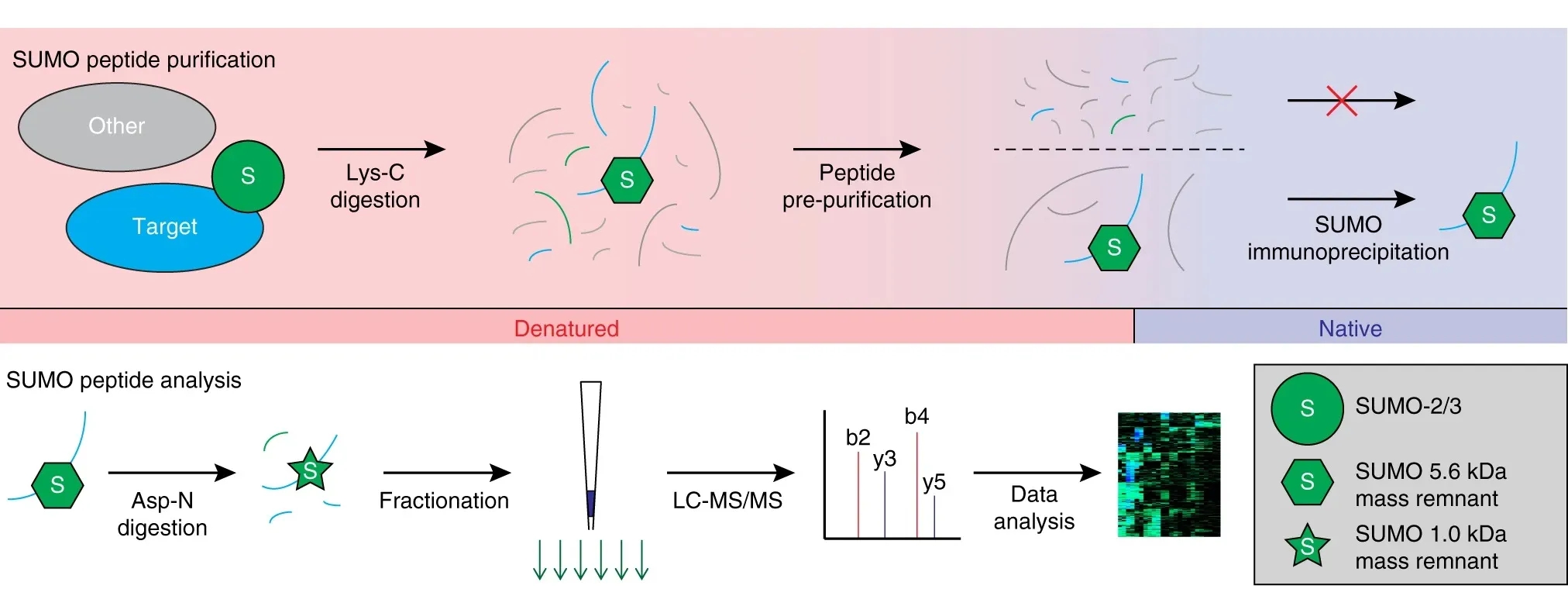
Hendriks, I. A. et al. Nat. Commun. 2018.
Figure 2. Workflow of Targeted SUMOylation Site Analysis
Why Choose MtoZ Biolabs?
1. Advanced LC-MS/MS Platform
High-resolution Orbitrap systems ensure accurate site identification and quantification of SUMOylated residues.
2. Optimized SUMO-Enrichment Strategy
Proprietary SUMO-remnant enrichment workflow ensures high specificity and recovery for site-level analysis.
3. Comprehensive Quantitative Capabilities
Flexible quantification methods provide reliable comparison of SUMOylation site abundance between samples.
4. Integrated Bioinformatics Analysis
Professional interpretation links modification sites to protein domains and functional pathways.
5. Experienced Proteomics Team
Expert scientists deliver detailed analytical reports with clear, actionable data summaries.
Applications of Targeted SUMOylation Site Analysis Service
✅ Mapping SUMOylation sites on regulatory or structural proteins
✅ Investigating SUMO-dependent mechanisms in DNA repair and chromatin organization
✅ Quantifying SUMOylation changes under stress or treatment conditions
✅ Studying SUMOylation-mediated control of transcriptional activity
✅ Characterizing interplay between SUMOylation and other PTMs
Start Your Project with MtoZ Biolabs
Partner with MtoZ Biolabs to achieve precise and reproducible SUMOylation site characterization for your specific proteins of interest. Our experienced team and advanced analytical platforms are ready to support your research from sample preparation to data interpretation.
Contact us today to discuss your experimental design or request a quote.
FAQ
Q1: What types of samples are suitable?
MtoZ Biolabs accepts diverse sample types including cell lysates, tissue extracts, and purified proteins. For optimal results, a minimum of 50 μg total protein is recommended per analysis.
Q2: What is the service general workflow?
Q3: What data formats are provided?
Clients receive a complete data package containing:
· Experimental report and workflow summary
· Raw LC-MS/MS data files
· Peptide and protein identification tables
· Site-level SUMOylation mapping and quantification results
· Quality control and statistical overview
All data are provided in standard formats (Excel, PDF, and instrument raw files) suitable for publication and further bioinformatics analysis.
Q4: How should I prepare my samples?
· Use freshly prepared or properly frozen lysates stored at −80°C.
· Avoid high-salt buffers or detergents such as SDS.
· Filter or centrifuge to remove debris before shipment.
For more information, please refer to Sample Submission Guidelines for Proteomics.
Related Services
How to order?







