Resources
Proteomics Databases
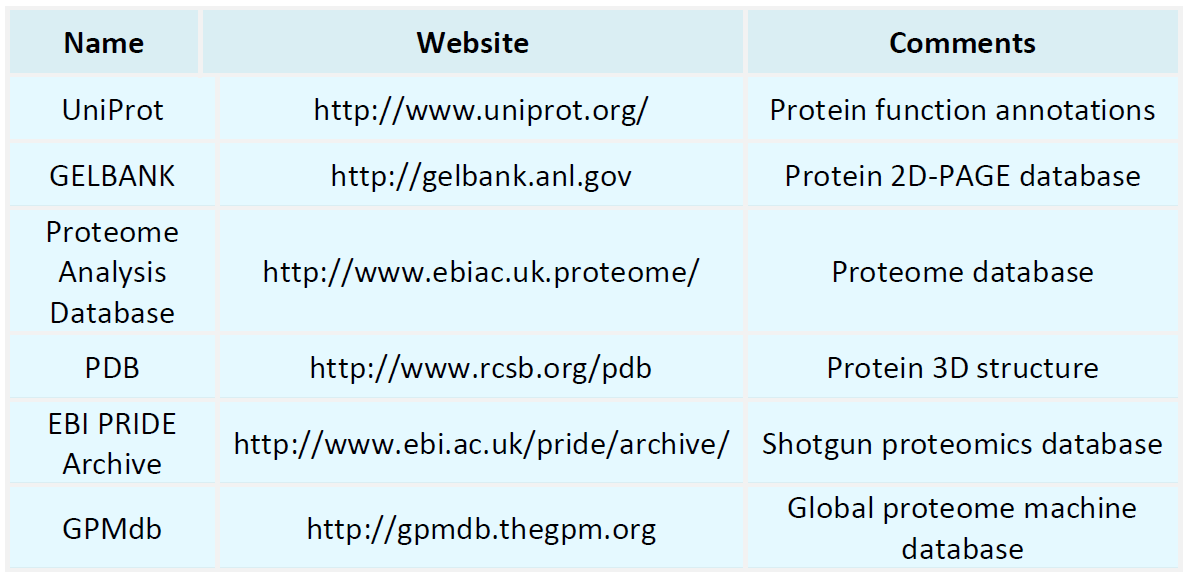
Metabolomics Databases
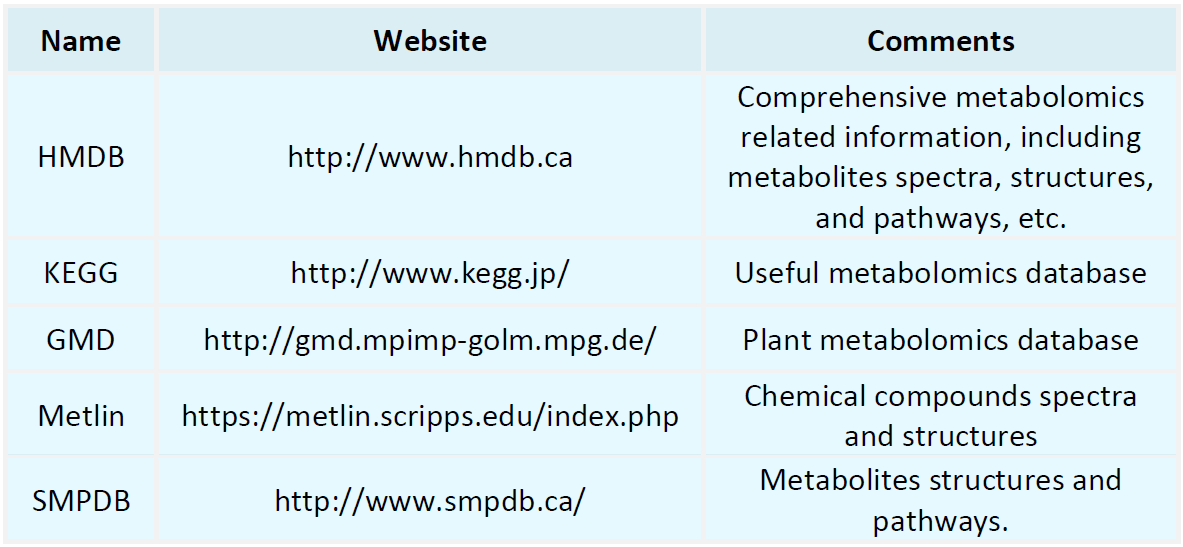
-
• Edman Degradation: Identifying Amino Acid Sequences for Peptides
Edman degradation is a chemical method used to determine the N-terminal amino acid sequence of peptides, such as proteins and peptide segments, in a sequential manner. This method was developed by Pehr Edman in the 1950s.
-
• Amino Acid Sequence Analysis and Its Applications
Amino acid sequence analysis is a fundamental field in protein research that involves the identification and interpretation of the linear sequence of amino acids in proteins. The amino acid sequence of a protein can provide important information about its structure, function, and evolution. Here are the applications of amino acid sequence analysis in protein research.
-
• Standard Protein Detection in Mass Spectrometry Analysis
Mass spectrometry is a powerful tool used in the field of proteomics for protein identification and quantification. The basic workflow for protein detection using mass spectrometry is outlined below.
-
• Mass Spectrometry: Method to Sequence Proteins
Mass spectrometry analysis has become the mainstream method for determining protein amino acid sequences. With appropriate preliminary preparation and mass spectrometry techniques, thousands of proteins can be identified and quantified from complex samples. The following are the basic steps for using mass spectrometry to analyze protein sequences.
-
• The Basic Process of Antibody Sequencing
Antibody sequencing is a technique used to determine the amino acid sequence of specific antibody molecules in monoclonal or polyclonal antibodies. This provides crucial information for understanding the specificity, activity, structure, and affinity of antibodies with antigens, and is important for antibody engineering, vaccine development, disease diagnosis, and therapeutic research.
-
• Guidelines for Protein Sequence Analysis Using the SMART Tool
"SMART" (Simple Modular Architecture Research Tool) is an online tool used to identify and analyze domains and functional sites in protein sequences. The basic steps to perform protein sequence analysis using SMART are as follows.
-
• Antibody Amino Acid Sequencing Basic Process
Antibody amino acid sequencing is performed to determine the amino acid sequence of the heavy and light chains of an antibody. This is typically done to address issues related to antibody specificity, activity, structure, and affinity for the antigen. The process of antibody amino acid sequencing generally involves the following steps.
-
• Protein Amino Acid Sequencing: Unveiling Key Traits in Protein
Protein amino acid sequencing is the process of determining the sequence of amino acids in a protein molecule. This process is crucial for understanding the structure and function of proteins, as protein function is often closely related to its amino acid sequence.
-
• Overview of Methods for Protein Amino Acid Sequence Detection
Protein amino acid sequence detection is performed to determine the primary structure of proteins, which refers to the linear arrangement of amino acids, as they are the building blocks of proteins and arranged in a specific linear sequence. Understanding this primary structure is similar to deciphering the genetic code of proteins and it holds significant importance in various scientific disciplines.
-
• Overview of Key Protein Structure Analysis Techniques
Protein function is directly determined by its structure, interactions with other proteins, and its location within cells, tissues, and organs. Large-scale studies of protein structure and function in proteomics have made it possible to identify protein biomarkers associated with specific disease states and provide potential targets for treatment.
How to order?







