Resources
Proteomics Databases
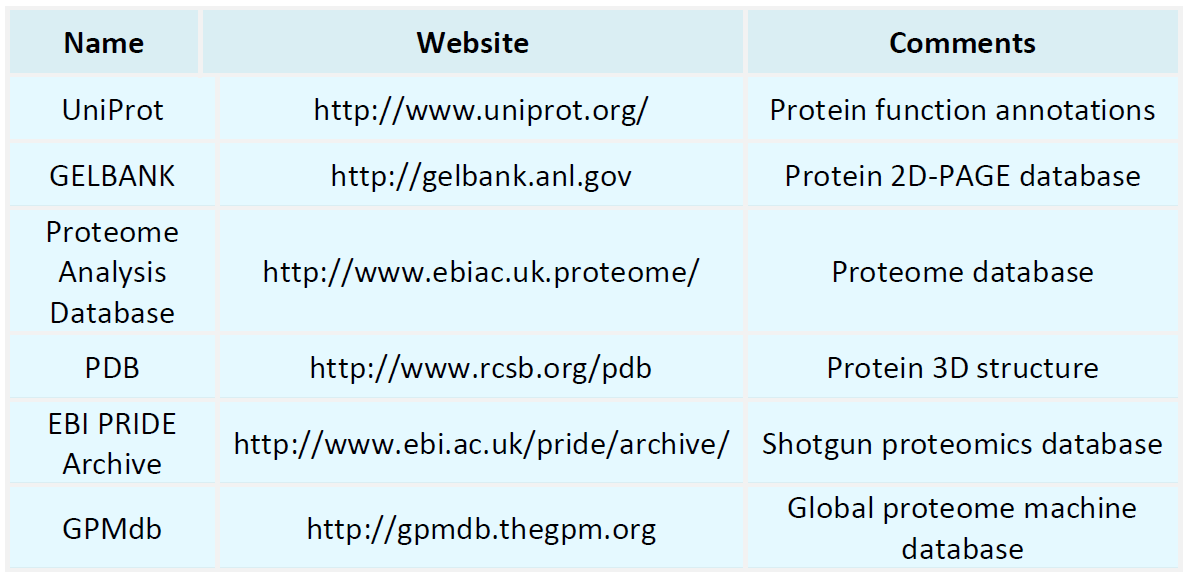
Metabolomics Databases
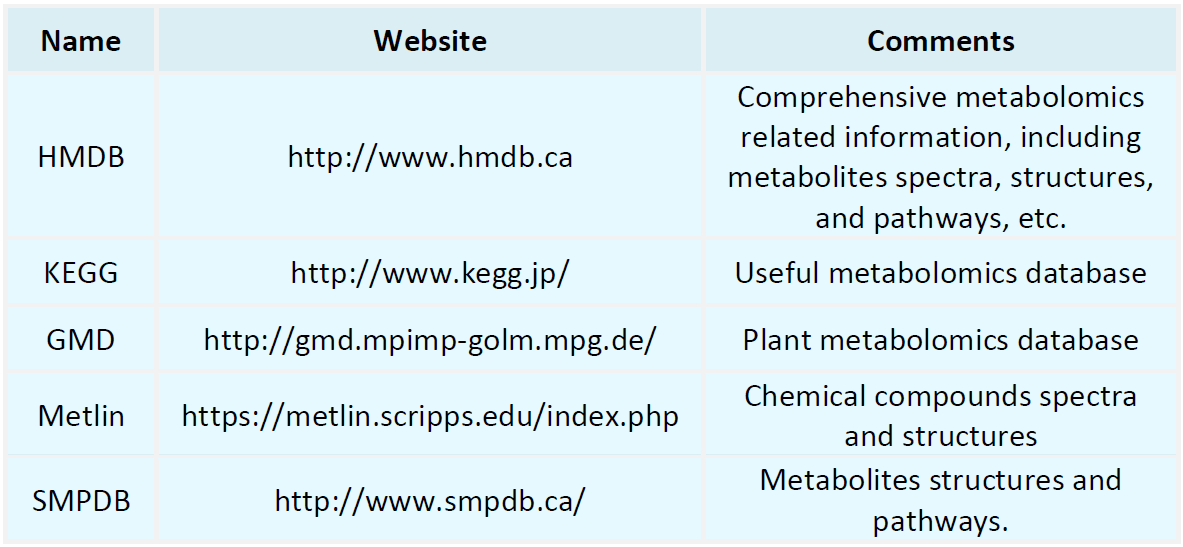
-
• Analysis of O-Glycans via Chemical Hydrolysis
O-glycans are an essential form of glycosylation in glycoproteins, playing crucial roles in regulating cellular communication and immune recognition. Studying the structure and function of O-glycans provides insights into these biological processes. Chemical hydrolysis is a commonly used technique to release and analyze O-glycans from glycoproteins.
-
• Analysis of N-Glycan Profiles Using MALDI-TOF-MS
N-glycans are a crucial modification of glycoproteins on the cell surface, involved in various biological processes such as protein folding, cell-cell communication, and immune response. Accurate analysis of N-glycan structures and compositions is essential for understanding their biological and medical roles. In recent years, matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF-MS) has become a powerful tool for studying N-glycan profiles
-
• Detection of O-Glycans by MALDI-TOF-MS
O-glycans, a common type of glycosylation modification, are widely present on the surface of proteins and play critical roles in various biological processes such as cell recognition, signal transduction, and immune responses. Due to the structural complexity and high heterogeneity of O-glycans, mass spectrometry-based methods have become indispensable tools for analyzing glycoproteins and studying glycosylation patterns.
-
• Detection of Glycosylation Sites and Glycoforms Based on LC-MS/MS
Glycosylation is a crucial post-translational modification that influences protein folding, stability, and function. Accurate detection of glycosylation sites and glycoforms is essential for studying biological processes, disease mechanisms, and drug development. Liquid chromatography-tandem mass spectrometry (LC-MS/MS) has become the primary method for detecting glycosylation sites and glycoforms due to its high sensitivity and resolution.
-
• Mechanism of Chemical Proteomics Analysis
Chemical proteomics is a research approach that combines chemical tools with mass spectrometry to study protein function, interactions, and modification states. The central mechanism involves using chemical probes, reactive compounds, or labeling molecules to identify proteins or their modification sites, followed by qualitative and quantitative analysis using mass spectrometry.
-
• Application of Chemical Proteomics Analysis
Chemical proteomics, a technique that utilizes chemical probes and targeted reactions to detect and functionally analyze proteome components, has found broad applications in the biomedical field. With advancements in analytical techniques, chemical proteomics has become a powerful tool for biological research and drug development. Its application spans from target identification to biomarker discovery, advancing research in precision medicine and drug development.
-
• Principle of Chemical Proteomics Analysis
Chemical proteomics is an emerging field that leverages the interaction between chemical probes and proteins to study protein function, structure, and activity within the cellular environment. By integrating chemical synthesis with proteomics techniques, it allows the identification, quantification, and functional analysis of proteins through specific chemical modifications or labeling.
-
• Workflow of Chemical Proteomics Analysis
Chemical proteomics is a technique that utilizes chemical methods to label, capture, or modify proteins in order to study their structure and function. This analytical technology has broad applications in drug discovery, biomarker identification, and protein function research. The workflow of chemical proteomics analysis typically involves several key steps, each of which is crucial for the success of the experiment. Below is a detailed description of the workflow of chemical proteomics analysis.
-
• Mechanism of AQUA in Protein Detection
Absolute Quantification of proteins (AQUA) is a key technology in proteomics, utilizing stable isotope-labeled peptides for precise protein quantification. Combined with mass spectrometry (MS), AQUA enables the absolute quantification of target proteins, providing crucial insights into biomarker discovery, drug target validation, and functional analysis of proteins.
-
• Workflow of AQUA in Quantitative Proteomics
AQUA (Absolute Quantification) is an isotope-labeled mass spectrometry-based technique used for the precise absolute quantification of specific peptides from target proteins. By incorporating known amounts of synthetic peptides as internal standards, AQUA provides absolute quantification of target peptides in a sample. The workflow of AQUA involves multiple critical steps, ranging from sample preparation to data processing and analysis, which ensures accuracy and reproducibility.
How to order?







