Integrated iTRAQ with LC-MS/MS Workflow for Quantitative Proteomics
-
Reporter group: Releases characteristic reporter ions upon MS/MS fragmentation for quantitative measurement.
-
Balance group: Adjusts the overall mass so that all labeled peptides remain isobaric and indistinguishable at the MS1 level.
-
Peptide reactive group: Facilitates covalent binding to peptides.
-
Investigation of disease mechanisms: Quantitative comparison across multiple groups enables identification of differentially expressed proteins. Integration with Gene Ontology (GO) and KEGG pathway enrichment analyses facilitates mechanistic interpretation.
-
Drug target discovery: Proteomic profiling before and after pharmacological intervention supports identification of candidate targets and potential biomarkers.
-
Biomarker discovery: In oncology and neurodegenerative disease research, iTRAQ enables detection of differentially expressed proteins within complex biological backgrounds.
-
Time-resolved dynamic studies: Suitable for developmental biology and time-course experiments, allowing precise characterization of temporal protein expression changes.
-
Optimization of sample preparation (tissue, body fluid, or cell lysis).
-
Proteolytic digestion and iTRAQ labeling.
-
High-pH reversed-phase fractionation to enhance proteome depth.
-
High-resolution LC-MS/MS analysis.
-
Differential protein identification and downstream bioinformatics analysis.
The advancement of proteomics depends on quantitative strategies that are high-throughput, accurate, and reproducible. Particularly when comparing protein expression across multiple biological states, such as disease models, pharmacological interventions, or developmental stages, stable isotope labeling approaches offer distinct advantages. Among these, the iTRAQ (Isobaric Tags for Relative and Absolute Quantitation)-LC-MS/MS-based quantitative workflow is widely adopted in quantitative proteomics owing to its multiplexing capability, high sensitivity, and strong data consistency.
Principle of iTRAQ Technology
iTRAQ is an isotopic chemical labeling strategy that enables simultaneous multiplexed quantification of up to eight, and in some implementations ten, samples within a single analytical run. Following proteolytic digestion, isotopic tags with identical total mass but distinct fragmentation properties are covalently attached to the N-terminus and lysine side chains of peptides. Each tag consists of three functional components:
At the MS1 stage, peptides labeled with different tags exhibit identical m/z values. Upon MS/MS fragmentation, tag-specific reporter ions with distinct masses are generated, enabling relative quantification across multiple samples.
Rationale for Choosing iTRAQ-LC-MS/MS Quantification
1. High-Throughput Multiplexed Analysis
iTRAQ supports multiplexed analysis of up to eight samples per experiment (iTRAQ 8-plex), substantially improving analytical efficiency. This feature is particularly advantageous for study designs involving time-course experiments, multiple treatment groups, or biological and technical replicates.
2. Enhanced Detection of Low-Abundance Proteins
The isobaric labeling strategy consolidates peptide signal intensity at the MS1 level, thereby improving the detectability of low-abundance peptides. This advantage is especially relevant for complex biological matrices, including serum and tissue homogenates.
3. Reduction of Inter-Sample Technical Variation
Following labeling, samples are pooled into a single mixture and analyzed in the same LC-MS/MS run. This approach minimizes batch effects introduced during sample preparation and instrumental analysis, thereby enhancing quantitative accuracy and reproducibility.
4. Compatibility with High-Resolution LC-MS/MS Platforms
iTRAQ labeling does not substantially alter peptide chromatographic behavior and is fully compatible with high-resolution LC-MS/MS systems, including Orbitrap and Triple TOF instruments. The workflow is well-established and technically robust.
Comparison of iTRAQ with Other Quantitative Strategies
When selecting a mass spectrometry-based quantification strategy, researchers often compare iTRAQ with TMT, SILAC, and label-free approaches. Each method has specific strengths and limitations depending on experimental context.
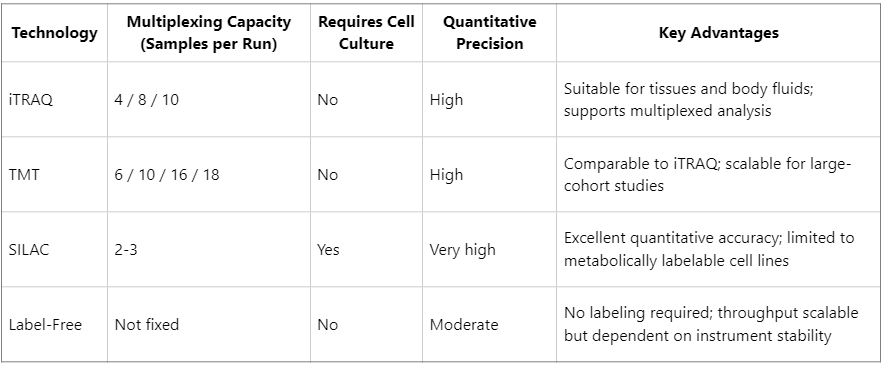
Collectively, iTRAQ is particularly suitable for experimental scenarios characterized by moderate sample numbers, non-cell culture specimens, and a requirement for high quantitative consistency, such as clinical tissues, animal models, and complex biological fluids.
Applications: From Differential Expression Profiling to Pathway Analysis
iTRAQ-LC-MS/MS has been extensively applied across diverse research areas:
iTRAQ Quantitative Workflow at MtoZ Biolabs
At MtoZ Biolabs, a standardized and reproducible iTRAQ-LC-MS/MS workflow has been established based on the high-resolution Orbitrap Fusion Lumos platform. The service pipeline includes:
Extensive experience in labeling efficiency assessment, quantitative bias control, and statistical analysis ensures methodological rigor and data reliability. This integrated workflow supports both clinical collaborative research and fundamental biological investigations through comprehensive iTRAQ-based quantitative proteomics services.
The iTRAQ-LC-MS/MS quantitative strategy remains a central approach in proteomics owing to its high throughput, reproducibility, and quantitative accuracy. Although alternative isobaric labeling technologies such as TMT have expanded multiplexing capacity in recent years, iTRAQ continues to represent a robust, reliable, and cost-balanced solution in appropriate research contexts. For studies investigating protein expression changes across multiple biological conditions, particularly comparative analyses involving tissues and body fluids, iTRAQ remains a well-validated and technically sound option. MtoZ Biolabs is committed to delivering high-standard, high-throughput proteomics solutions to support advanced quantitative research.
MtoZ Biolabs, an integrated chromatography and mass spectrometry (MS) services provider.
Related Services
How to order?







