What Sample Preparation Methods Are Suitable for FFPE Proteomics?
-
Extensive crosslinking: Formaldehyde forms covalent crosslinks with protein side chains, reducing enzymatic digestion efficiency
-
Risk of protein degradation: Long-term storage may result in protein modifications or partial degradation
-
Limited sample availability: This is particularly evident in biopsy specimens or microdissected tissue samples
-
Efficient deparaffinization
-
Effective crosslink reversal
-
Mass spectrometry-compatible protein purification and enzymatic digestion
-
Xylene-mediated deparaffinization
-
Gradient ethanol rehydration
-
Protein solubilization using SDS lysis buffer
-
High-temperature incubation at 95–100 °C to reverse cross-links
-
Trypsin digestion
-
Well-established protocol
-
Suitable for routine batch processing
-
Relatively high protein recovery
-
SDS interferes with mass spectrometry and must be removed during downstream processing (e.g., FASP or SP3)
-
Less suitable for low-input samples
- Clinical tissue blocks with relatively sufficient protein amounts (>50 μg)
-
Proteins are first extracted using SDS lysis and subsequently loaded onto an ultrafiltration membrane.
-
SDS removal and buffer exchange are performed on the membrane.
-
Enzymatic digestion is carried out directly on the filter.
-
Efficient removal of SDS
-
Improved enzymatic digestion efficiency
-
Good data reproducibility
-
Relatively higher sample loss
-
Less suitable for ultra-low input samples
- Routine proteomics analysis and preprocessing for label-based quantification strategies such as TMT
-
Proteins are captured by magnetic beads under high concentrations of organic solvents.
-
Contaminants are removed during washing steps.
-
Enzymatic digestion is performed directly on the beads.
-
Highly suitable for low-input samples (even <1 μg)
-
Low sample loss
-
Automation-compatible workflow
-
Compatible with TMT and DIA strategies
-
Sensitive to operational parameters
-
Magnetic bead ratios require optimization
- Microdissected tissues, biopsy samples, and laser capture microdissection specimens
-
Proteins are acid-precipitated onto a quartz fiber matrix.
-
Rapid SDS removal
-
On-column enzymatic digestion
-
Simple workflow
-
Suitable for moderate sample amounts
-
Good reproducibility
- Standardized experimental workflows and large-scale batch analyses
-
Tissue sectioning → deparaffinization → microscopic localization
-
Laser-based precise excision of the target region
-
Low-input protein extraction followed by SP3 preparation
-
High spatial resolution
-
Enables studies of tumor heterogeneity
-
Technically demanding
-
Extremely limited sample amounts require highly optimized workflows.
-
Optimization of crosslink reversal temperature and duration (typically 95 °C for 1–2 hours)
-
pH control during reversal (Tris buffer facilitates formaldehyde reversal)
-
Extended enzymatic digestion time to improve identification depth
-
Integration with high-resolution mass spectrometry platforms
-
Tumor biomarker discovery
-
Prognostic molecular subtyping
-
Prediction of immunotherapy responses
-
Retrospective cohort validation
-
Support paraffin sections, tissue blocks, and microdissected samples.
-
Provide multiple preparation strategies including SP3, S-Trap, and FASP.
-
Ensure compatibility with DIA and TMT quantitative workflows.
-
Deliver deep and highly reproducible proteomics datasets.
In clinical research and translational medicine, Formalin-Fixed Paraffin-Embedded (FFPE) tissue samples represent one of the most widely available and longest-preserved types of biological materials. Large FFPE archives have been accumulated in hospitals and pathology departments worldwide, encompassing diverse specimen types such as tumors, inflammatory diseases, and rare disorders. These collections provide unique opportunities for retrospective cohort studies. However, FFPE proteomics is far more complex than routine protein extraction. The formalin fixation process introduces methylene bridge crosslinking, which alters protein conformation and decreases protein solubility, thereby posing significant challenges for downstream mass spectrometry analysis. Consequently, selecting an appropriate sample preparation strategy is critical for obtaining high-quality FFPE proteomics data.
Core Challenges of FFPE Proteomics
Before discussing specific methodologies, three major technical challenges associated with FFPE samples should be clarified:
Therefore, an ideal FFPE preparation workflow should include:
Mainstream FFPE Sample Preparation Methods
1. SDS Lysis Combined with High-Temperature Crosslink Reversal (Classical Method)
(1) Technical Principle
(2) Advantages
(3) Limitations
(4) Applicable Scenarios
2. FASP Method (Filter-Aided Sample Preparation)
FASP is one of the most widely used strategies in FFPE proteomics.
(1) Technical Principle
(2) Advantages
(3) Limitations
(4) Applicable Scenarios
3. SP3 Method (Single-Pot Solid-Phase-Enhanced Sample Preparation)
(1) Technical Principle
(2) Advantages
(3) Limitations
(4) Applicable Scenarios
4. S-Trap Method
(1) Technical Features
(2) Advantages
(3) Applicable Scenarios
5. Laser Capture Microdissection (LCM) Combined with Low-Input Preparation
(1) Technical Workflow
(2) Advantages
(3) Limitations
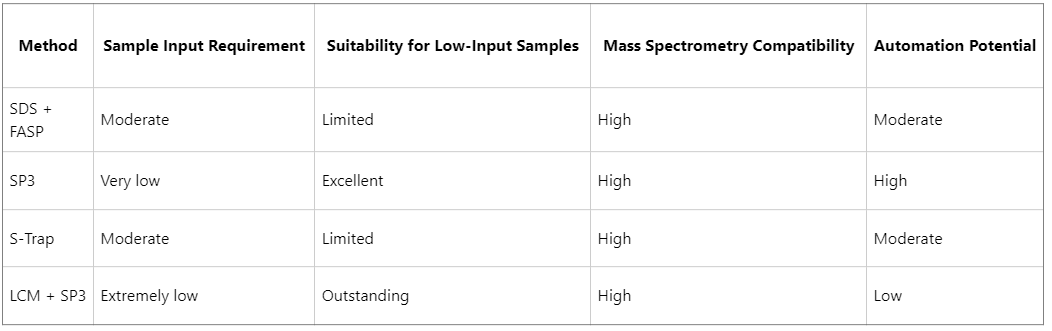
Comparative Summary of Different Methods
Key Optimization Considerations in FFPE Proteomics
Modern FFPE proteomics studies frequently utilize high-resolution Orbitrap or timsTOF mass spectrometry systems, enabling deep proteome coverage exceeding 5,000 identified proteins.
Application Prospects of FFPE Proteomics
FFPE samples have been widely applied in:
In the era of precision medicine, FFPE proteomics is becoming an essential bridge linking clinical pathology with molecular mechanisms.
Professional Technical Support: High-Quality FFPE Proteomics Solutions
In real-world studies, FFPE samples often vary significantly in preservation conditions and crosslinking levels. Therefore, optimized workflows frequently need to be customized according to specific sample types.
Leveraging advanced high-resolution mass spectrometry platforms and a well-established FFPE sample preparation system, MtoZ Biolabs is able to:
Through multiple clinical collaboration projects, large-scale FFPE cohort analyses have been successfully completed, facilitating the discovery of potential biomarkers.
Selecting an appropriate preparation strategy (SDS, FASP, SP3, S-Trap, or LCM-integrated workflows) requires comprehensive consideration of sample input, research objectives, and analytical strategies. If you are planning FFPE proteomics studies or encountering challenges such as limited sample availability or insufficient proteome coverage, you are welcome to consult the technical team at MtoZ Biolabs. We provide professional and reliable mass spectrometry solutions to support your research projects.
MtoZ Biolabs, an integrated chromatography and mass spectrometry (MS) services provider.
Related Services
How to order?







