What Are the Challenges of Proteomics on FFPE Samples?
-
Crosslinking alters protein spatial conformation, thereby hindering efficient enzymatic digestion by proteases such as trypsin.
-
It induces alterations in protein post-translational modifications (PTMs), including formaldehyde-induced methylation, thereby affecting quantitative accuracy.
-
Protein solubility and extraction efficiency are reduced, with particularly pronounced effects on membrane proteins and low-abundance proteins.
-
Deparaffinization and rehydration: Incomplete removal of paraffin can severely suppress ionization efficiency in mass spectrometry.
-
Antigen retrieval: Conditions such as high temperature/high pressure or enzymatic digestion must be optimized to reverse crosslinking.
-
Tissue lysis: Fixed tissues exhibit increased structural rigidity, necessitating the use of strong lysis buffers or mechanical homogenization to enhance extraction efficiency.
-
Variability in the extent of protein degradation among samples.
-
Differences in fixation conditions across batches affecting protein extraction efficiency and modification states.
-
Accumulation of non-specific modifications, such as oxidation and carbonylation, during long-term storage.
-
Difficulty in detecting low-abundance proteins, including key regulators of cellular signaling pathways.
-
Quantitative measurements being affected by modification-induced interference, necessitating careful consideration of antibody specificity in immunoaffinity enrichment-MS workflows.
-
Reduced labeling efficiency in tag-based quantification strategies (e.g., TMT/iTRAQ).
Among clinical sample resources, formalin-fixed paraffin-embedded (FFPE, Formalin-Fixed Paraffin-Embedded) tissues, owing to their advantages such as long-term preservation, abundant availability, and broad coverage of pathological types, represent a highly valuable resource for proteomics research. They are particularly valuable in disease biomarker discovery, tissue-specific protein studies, and retrospective cohort analyses. However, FFPE samples also present numerous technical challenges during proteomic analysis.
Formaldehyde Crosslinking Induces Protein Denaturation and Modifications
The core preservation mechanism of FFPE samples is formaldehyde crosslinking (formalin fixation), which stabilizes protein molecules within tissues through covalent bonding. However, this process inevitably introduces several issues:
These chemical modifications are often reflected in mass spectrometry as reduced peptide identification rates and increased spectral complexity.
Challenges in Sample Preparation Introduced by Paraffin Embedding
While paraffin embedding preserves tissue architecture, several critical steps are required prior to mass spectrometry analysis:
These procedures impose stringent requirements on experimental stability and reproducibility, and thus require careful control of the workflow.
High Sample Heterogeneity and Pronounced Batch Effects
FFPE samples are often derived from clinical tissues spanning different centers, pathological conditions, and storage durations, leading to:
Collectively, these factors exacerbate batch effects and increase the complexity of data interpretation, posing challenges for cross-cohort comparative analyses.
Limitations in the Accuracy and Sensitivity of Protein Quantification
Protein yield in FFPE samples is often reduced due to degradation and chemical modifications, resulting in:
Although high-sensitivity mass spectrometers, such as Orbitrap Eclipse and timsTOF Pro, have substantially improved detection capabilities, optimal performance still depends on well-optimized sample preparation strategies.
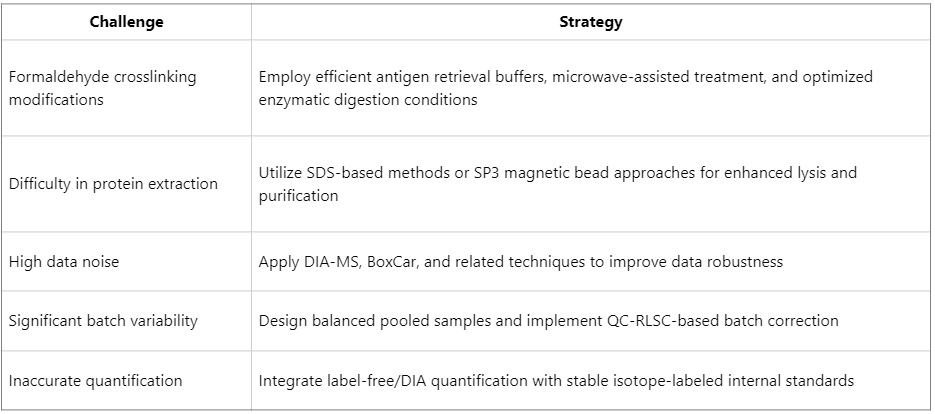
Overview of Solutions and Emerging Strategies
Despite these challenges, FFPE proteomics is not infeasible. Recent advances in sample processing and mass spectrometry technologies have enabled the development of strategies to address these limitations:
At MtoZ Biolabs, we have established a dedicated proteomics workflow for FFPE samples, encompassing efficient deparaffinization and antigen retrieval, low-input protein enrichment strategies, and a DIA-based quantitative platform. This workflow has been successfully applied to multiple studies in cancer and neurodegenerative diseases.
Given their clinical relevance and accessibility, FFPE tissue specimens are becoming increasingly important in proteomics research. Although chemical fixation and processing introduce substantial technical challenges, high-quality proteomic data can be achieved through appropriate experimental design, optimized sample preparation, and advanced mass spectrometry platforms. Looking forward, continued standardization of sample processing, improvements in mass spectrometry sensitivity, and the integration of AI-assisted data analysis are expected to further expand the role of FFPE proteomics in precision medicine and biomarker development. For more information on FFPE-based mass spectrometry solutions, please contact MtoZ Biolabs for customized project consultation and technical support.
MtoZ Biolabs, an integrated chromatography and mass spectrometry (MS) services provider.
Related Services
How to order?







