Step-by-Step Protocol for Conducting ABPP Experiments
- Extract proteins from cells or tissues while preserving enzymatic activity
- Incubate the sample with the probe for 30-60 min
- Terminate the reaction and proceed with downstream processing
- Add the probe directly to live cells
- Rapidly lyse the sample after the reaction is complete
- Capture enzyme activity profiles under near-physiological conditions
- Screening and synthesis of multiple activity-based probes, including customized probe design
- End-to-end experimental workflow support, from sample preparation to enrichment and proteolytic digestion
- High-resolution mass spectrometric analysis and data processing
- Differential enzyme activity analysis, pathway annotation, and visualized data output
Unlike conventional proteomics approaches, ABPP (Activity-Based Protein Profiling) not only identifies protein expression, but also captures protein activity states. It is particularly well suited for functionally important enzyme classes with complex regulatory behavior, such as proteases, phosphatases, and deacylases. How, then, can a complete ABPP experiment be systematically designed and implemented? From probe selection to mass spectrometric analysis, and from sample preparation to data interpretation, this article provides a step-by-step breakdown of the full ABPP workflow and offers a clear and efficient practical guide for your research.
The Complete Workflow Of An ABPP Experiment
1. Define The Research Objective And Enzyme Family
The first step in an ABPP experiment is to clearly define the research objective: which enzyme class is to be analyzed, and under which biological model or treatment condition will the experiment be performed? Common objectives include:
(1) Screening proteases with aberrant activity in tumors
(2) Evaluating the effects of candidate drugs on target enzymes
(3) Analyzing the regulatory patterns of active enzymes in inflammatory or apoptotic signaling pathways
Different enzyme classes require specifically designed or selected ABPs (Activity-Based Probes). Therefore, defining the target enzyme class determines the subsequent probe strategy and downstream data analysis framework.
2. Select Or Customize Activity-Based Probes (ABPs)
ABPs are the core tools underlying successful ABPP experiments. Structurally, they generally consist of the following three components:
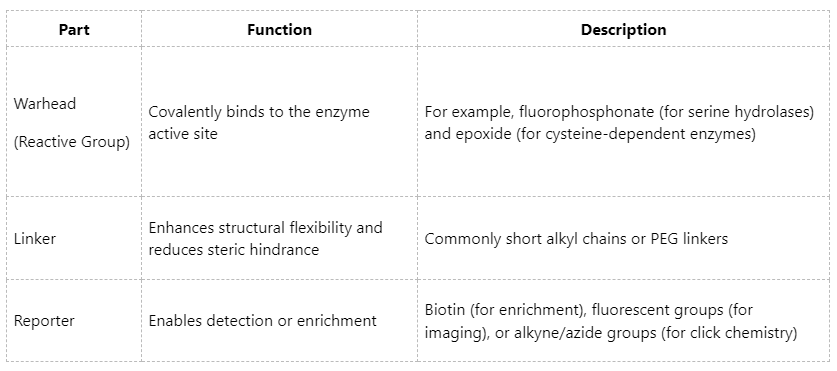
3. Probe Labeling Of ABPP Samples
Probe-labeling experiments are generally performed in two main modes:
(1) Cell Lysate Labeling (In Vitro)
This approach is suitable for enzyme screening and studies of drug mechanism of action.
(2) Live-Cell Or Tissue Labeling (In Situ/In Vivo)
This approach more closely reflects the physiological state, although probe permeability and toxicity must be carefully considered.
4. Enrichment And Proteolytic Digestion
Once labeling is complete, the next step is to enrich the labeled active enzymes from the complex biological background:
(1) If biotin labeling is used, streptavidin magnetic beads can be applied for affinity purification
(2) If click-functionalized probes are used, a click reaction is first performed, followed by conjugation to an enrichment tag such as biotin-azide
After enrichment, standard proteolytic digestion is performed using trypsin or other proteases to generate peptides for subsequent mass spectrometric analysis.
5. LC-MS/MS Analysis
After sample preparation, the workflow enters the stage of high-resolution mass spectrometric analysis. The following strategies are commonly recommended:
(1) Instrument platform: Orbitrap Exploris 480 or similar systems, suitable for high-throughput, high-accuracy analysis
(2) Separation mode: nanoscale reversed-phase liquid chromatography (nanoLC), which improves peptide separation efficiency
(3) Acquisition mode: DDA (data-dependent acquisition) or DIA (data-independent acquisition)
(4) Quantification strategy: Label-free, TMT, iTRAQ, and other quantitative approaches are all applicable
The raw data generated at this stage provide the core basis for subsequent activity profiling.
6. Data Analysis And Activity Profile Generation
The ultimate goal of ABPP is to generate an activity-based proteomic profile. The data analysis workflow mainly includes:
(1) Peptide Identification And Quantification
Use specialized software, such as MaxQuant or Proteome Discoverer, for peptide identification and quantification.
(2) Specificity Filtering
Retain only successfully labeled proteins captured by the probe, while removing background signals.
(3) Activity Change Analysis
Compare differences in enzyme activity levels between treatment groups, such as drug-treated versus control groups. Log2 fold change and FDR are commonly used as screening criteria.
(4) Functional Annotation And Pathway Enrichment
Integrate GO, KEGG, and Reactome databases to functionally classify highly active or differentially active enzymes and perform pathway enrichment analysis, thereby extracting biological significance.
(5) Data Visualization
Generate protease activity heatmaps, molecular interaction networks, pathway enrichment plots, and other visual outputs to support manuscript preparation or patent applications.
FAQ
Q1: What Types Of Proteins Are Suitable For ABPP?
ABPP is primarily used for enzymes that contain reactive active sites and can be covalently modified by probes, such as proteases, phosphatases, and ubiquitin-related enzymes.
Q2: How Can Probe Non-Specificity Be Addressed?
Non-specific background can be reduced by including a blank control group and a competitive inhibition group, while further optimization can be achieved by adjusting probe concentration and reaction time.
Q3: How Can Data Reproducibility Be Ensured?
At least three biological replicates are recommended. Quantitative strategies such as TMT can also be used to reduce batch effects.
MtoZ Biolabs: A One-Stop End-To-End ABPP Service Platform
To reduce the technical barriers associated with ABPP and improve experimental robustness, MtoZ Biolabs provides the following services:
Leveraging an advanced Orbitrap platform and a functional proteomics analysis framework, MtoZ Biolabs supports efficient progression from activity profiling to target discovery.
The value of ABPP lies in the fact that it reveals not only whether a protein is present, but also which proteins remain functionally active and are truly participating in biological processes. Mastering ABPP is not merely mastering a methodology; it is gaining a powerful means to understand protein functional states and enable precise target discovery. MtoZ Biolabs welcomes inquiries regarding professional training and project support for ABPP workflows, helping researchers move from expression-level analysis to function-oriented investigation.
MtoZ Biolabs, an integrated chromatography and mass spectrometry (MS) services provider.
Related Services
How to order?







