Single B-cell Screening and Antibody Research Development
Monoclonal antibodies (mAbs) are monovalent proteins expressed by B cells, capable of precisely targeting 3D epitopes, which makes it a compelling therapeutic option. Over 70 monoclonal antibody drugs have received approval. Monoclonal antibody drugs dramatically transform modern medicine by offering targeted treatments for a variety of diseases. These antibodies specifically activate immune pathways to combat cancer, suppress autoimmunity and excessive immune responses in transplantation to relieve cardiovascular diseases. Additionally, mAbs facilitate research into human B cell repertoires, the behavior of antibody-secreting cells post-infection and vaccination, response of memory B cells (MBC), and the response of pathogenic B cells in autoimmune diseases.
Single B-cell receptor (BCR) cloning rapidly produces antigen-specific mAbs within weeks. mAbs are generated through the pairing of B cell-derived heavy (VH) and light chains (VL). Initially, mAbs were produced by hybridoma cell forming by fusing a single B cell with an immortalized, immunoglobulin-deficient myeloma cell. Advanced techniques such as single BCR cloning and phage display libraries have now largely replaced hybridoma methods, becoming the standard for producing human mAbs. Single BCR cloning efficiently generates numerous antigen-specific mAbs quickly. In contrast, phage display libraries, while screening thousands of antigen-specific mAbs, typically yield only a few low-affinity antigen-specific outcomes. Moreover, these libraries predominantly produce mAbs through the random pairing of VH and VL isolated from from naïve B cells and MBCs, and do not adequately reflect B cell responses during infections, vaccinations, or autoimmune conditions. Consequently, single BCR cloning offers a effective, reliable, and fast approach to investigating B cell specificity across diverse disease scenarios.
In personalized medicine, a potent mAb production technique is vital for advancing biomedical and immunotherapy research. The monoclonal approach allows for in-depth studies of B cell biology, encompassing B cell development, antibody reactions to infections or vaccinations, and autoimmune diseases.
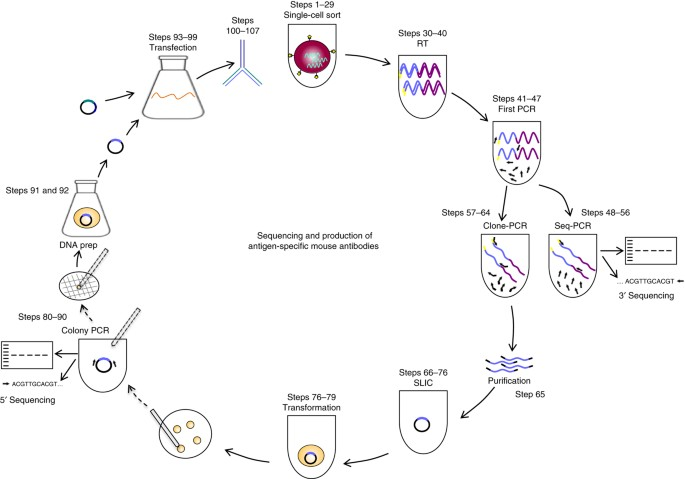
Figure 1. Schematic Diagram Illustrating the Process from Antigen-Specific Memory B-Cell Single-Cell Sorting to Antibody Purification [4]
1. Identification and Isolation of Single B-Cells
Depending on the application, single B-cell isolation can be performed either randomly or in an antigen-specific manner from peripheral blood or lymphoid tissues (bone marrow). Techniques for random B-cell isolation include micromanipulation for cell picking, laser capture microdissection, and fluorescence-activated cell sorting (FACS). For antigen-specific selection, magnetic beads coated with antigens and fluorescently labeled antigens are employed. Furthermore, a microarray chip system has been developed for the high-throughput screening (HTS) of cells that secrete mAb.
The primary advantage of using FACS in this context is its ability to distinctly categorize the developmental and differentiation stages of the cells targeted for sorting, based on their specific cell surface marker expression patterns. While B cells at any stage can be sorted, class-switched memory B cells and antibody-secreting cells (ASCs, i.e., plasmablasts and plasma cells) are especially targeted for obtaining high-affinity mAb because they possess somatically mutated BCR. Unlike these, vaccine-specific ASCs can be sorted without antigen labeling post-immunization. To efficiently isolate specific mAb, it is crucial to assess the donor's immune response, such as by determining the frequency of ASCs in peripheral blood using the enzyme-linked immunospot assay (ELISPOT) prior to single-cell isolation.
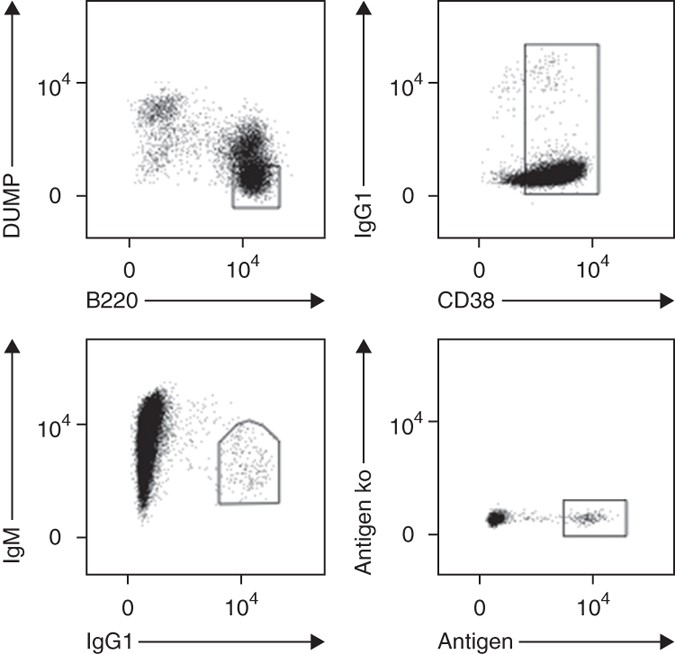
Figure 2. Gating Strategies for the Identification of Antigen-Specific Memory B Cells[4]
Currently, several high-throughput techniques allow for the effective identification of B cells with desired specificities prior to Ig gene cloning. (1) A soft lithography-based microarray method involves the individual deposition of polyclonal B cells onto a microwell array chip. Following this, a protein array is used to screen for B cells that produce specific antibodies. These cells are then manually extracted from the wells for further analysis. (2) immunospot array assay on a chip (ISAAC) places individual ASC on a microwell chip and detects the secreted antibodies by capturing them on the surface surrounding the wells. These techniques offer numerous advantages, such as the ability to rapidly and early identify cells with specific high-affinity antibodies from a polyclonal mixture, and simplify screening of various antigens to isolate different clones with diverse specificities.
2. Single-Cell Ig Gene Transcript Amplification, Sequencing, and Cloning
Constructing complementary DNA (cDNA) from a single B cell provides an unbiased approach for the simultaneous analysis of expressed IgH and IgL chain genes. Typically, this synthesis of single-cell cDNA occurs in the original equipment designed for cell deposition and lysis, such as a 96-well plate. This setup ensures the convenient processing of numerous samples while minimizing cross-contamination risks. Notably, various B cell types exhibit different quantities of specific Ig gene mRNA transcripts. For instance, ASCs, compared to memory B cells, possess a significantly higher number of Ig gene transcripts, thereby facilitating the specific amplification of these genes. Generally, full-length Ig gene transcripts are amplified using nested or semi-nested RT-PCR, where RT acts as the initial step of the PCR reaction. A mixture of forward primers, complementary to the leader sequences of the respective IgH and IgL V genes, and a specific single reverse primer for the constant region sequences, are employed. If necessary, such as during isotype-independent cell sorting, mixed reverse primers are used to amplify IgH chains featuring various constant regions. In the subsequent round, nested primers or primer mixtures enhance the sensitivity and specificity. Moreover, restriction sites crucial for subsequent cloning steps are integrated into the amplicons. Alternatively, linear expression cassettes are assembled and directly transfected into mammalian cells for the in vitro expression of amplified Ig genes. Methods to connect rearranged IgH and IgL chain genes during amplification have been explored. Research has led to the development of a multiplex single-cell RT-PCR method that joins PCR-amplified IgH and IgL chain products during PCR through an overlapping extension step. The resulting linked DNA fragments are then cloned into suitable expression vectors for mAb production.
Despite its straightforward nature, the single-cell RT-PCR method is limited by the design set of the used forward primers, and not every scheme successfully amplifies all functional IgH and IgL V genes. The "5' rapid amplification of cDNA ends" (5'-RACE) technique permits the amplification of the unknown 5' ends of mRNA. Since the standard 5' RACE technique is generally unsuitable for synthesizing cDNA from single cells due to the typically low mRNA amounts, improved schemes have demonstrated their effectiveness in isolating Ig genes from single B cells with high efficiency.
Irrespective of the chosen Ig gene amplification strategy, transcription information encoding antibody specificity from single-cell Ig genes is subsequently sequenced. Various databases, such as the NCBI's IgBLAST search engine (http://www.ncbi.nlm.nih.gov/igblast/), facilitate the easy identification and analysis of rearranged V, D, and J gene segments for mutations, insertions, and deletions.
3. Antibody Reactivity Screening
It is essential to determine the reactive properties and biophysical characteristics of antibodies or their fragments, expressing, purifying, and testing the proteins in various assays. Notably, this screening can be performed in a cell-based microarray system prior to Ig gene cloning, offering a streamlined approach. Nevertheless, for thorough analysis of the antibodies, molecular cloning of the Ig gene is crucial due to the significant protein quantities required. Common expression systems include bacterial systems such as Escherichia coli, which typically express cloned Ig genes as antigen-binding fragments (Fabs), and mammalian cell systems like HEK 293 or CHO cells, where the complete IgG form is expressed.
Analysis Workflow
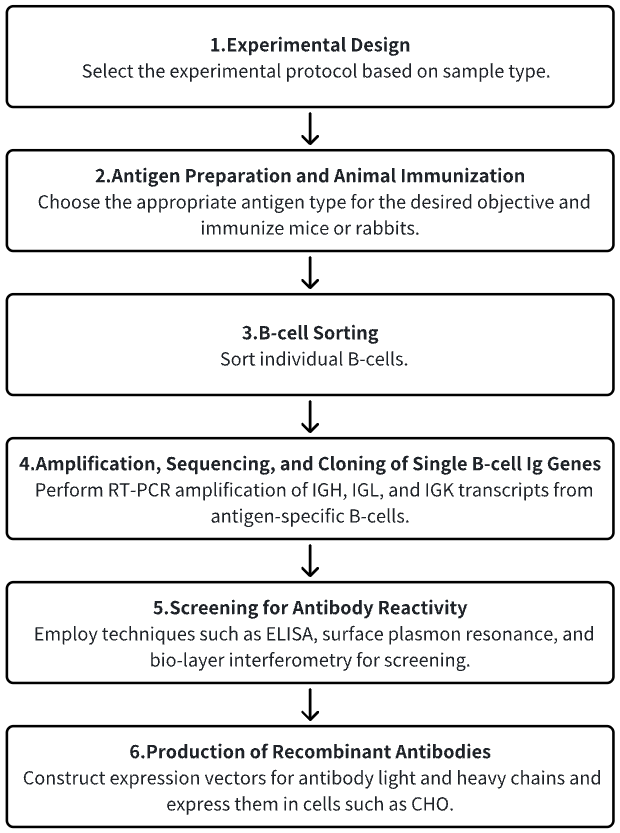
1. Definition of the Experimental Procedure Based on Needs
2. Animal Immunization
3. Sorting of Single B-Cells
4. Amplification, Sequencing, and Cloning of Single-Cell Ig Gene Transcripts
5. Screening for Antibody Reactivity
6. Production of Recombinant Antibodies on a Large Scale
Service Advantages
1. Monoclonal Antibody Preparation Across Multiple Species
2. Comprehensive Services for Single B-Cell Sorting and Screening
3. High-Accuracy Antibody Gene Sequencing
4. Efficient Construction and Expression of Vectors for Antibody Gene
Example Results
1. Recombinant Monoclonal Antibody Production from Immunized Mice and Rabbits through Flow Cytometry and Antigen-Specific IgG+ Memory B Cell Sorting
The single B cell screening strategy circumvents hybridoma fusion and combinatorial display, establishing itself as a crucial method for efficiently sampling natural Ab repertoire of immunized animals and humans. Various techniques are employed to assay different B-cell subset, providing a highly attractive approach for generating numerous and various high-quality Abs. The ability to produce multiple Abs and discover rare B cell clones that secrete IgGs with unique and desirable characteristics facilitates the identification of molecules suitable for development into therapeutic agents or research reagents. A multiparameter flow cytometry single-cell sorting technique was described for producing antigen-specific recombinant mAbs from individual IgG+ memory B cells. B cells from mouse spleen cells and rabbit peripheral blood mononuclear cells (PBMCs) of immunized animals were used as sources. Reagents staining both B cells and other non-targeted cell types effectively identified class-switched IgG+ memory B cells. By using two distinct fluorophores to stain for antigens, antigen-specific B cells-those binding to both antigen conjugates (double-positive)-were identifiable. Following this, cells were typically sorted at a rate of one cell per well into 96-well plates containing reverse transcription reaction mix using FACS. cDNA was then synthesized, followed by PCR amplification of cognate heavy and light chain variable region genes to create transcriptionally-active PCR (TAP) fragments. These linear expression cassettes were directly used for transfection in mammalian cells to produce recombinant antibodies, which were then subjected to further testing. The study successfully produced antigen-specific recombinant antibodies from both the rabbit and mouse IgG memory B cells subset within a week, including an anti-tumor necrosis factor receptor 2 (TNFR2) blocking antibody from mice with a 90 pM affinity.
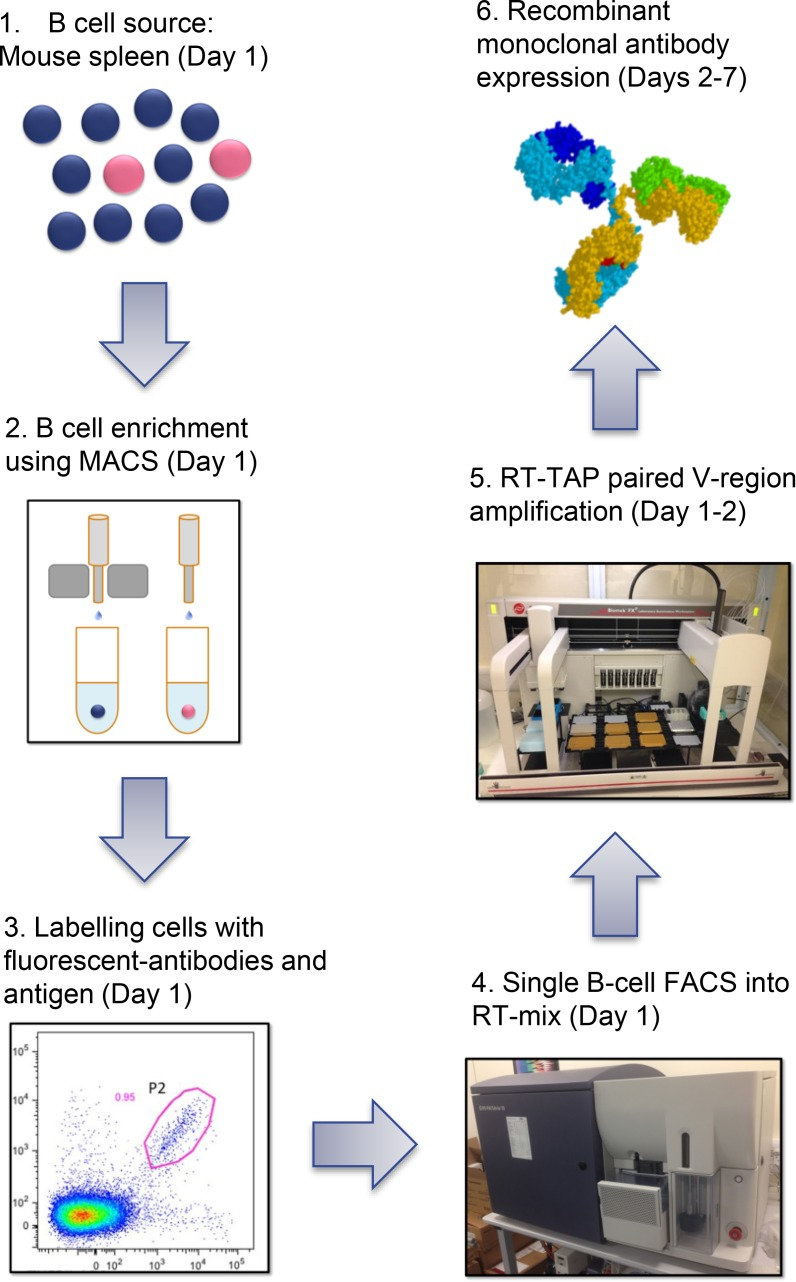
Figure 3. Diagram of Single B-Cell Sorting Approach for Antibody Discovery in Immunized Mice [5]
2. Massively Parallel Single-Cell B-Cell Receptor Sequencing for Rapid Identification of Antigen-Reactive Antibodies
The large-scale extraction of full-length variable regions for both heavy and light chains from individual B cells continues to pose significant challenges. Utilizing high-throughput single-cell B-cell receptor sequencing (scBCR-seq), studies have successfully obtained accurately paired full-length variable regions in a massively parallel fashion. Over 250,000 B cells from repertoires of rats, mice, and humans were sequenced to analyze their lineage and expansion. Moreover, rats were immunized with ovalbumin, and antigen-reactive B cells were analyzed from the lymph nodes of these immunized animals. The scBCR-seq data successfully recovered 81% (n = 56/69) of the B-cell lineages identified from the hybridomas produced from the same set of B cells subjected to scBCR-seq. Notably, scBCR-seq identified an additional 710 candidate lineages not restored as hybridomas. Researchers synthesized, expressed, and evaluated 93 clones from these identified lineages, discovering that 99% (n = 92/93) of the clones were antigen-reactive. These findings confirmed scBCR-seq as an effective tool for discovering antibodies.
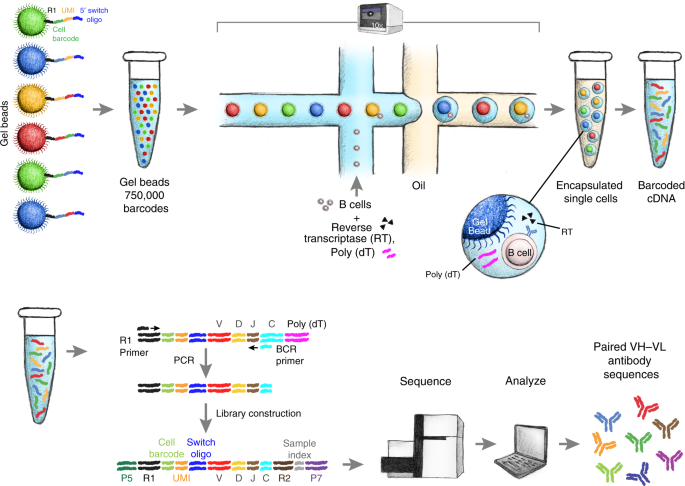
Figure 4. Schematic of Single B-Cell Capture, Library Construction, and Sequencing [6]
3. Single-Cell Sorting and Antibody Cloning of HBsAg-Binding Memory B Cells Derived from Human Peripheral Blood Mononuclear Cells
The isolation of human antibodies with inherently paired heavy and light chains is essential for understanding human immune responses to antibodies. A methodology has been proposed for Ab cloning from individual human memory B cells that recognized the Hepatitis B virus (HBV) S antigen (HBsAg). A dual-fluorescent-dye labeling approach enhanced the specificity of sorting against HBsAg, whereas staining with non-related proteins helped exclude non-specific B cells. This methodology was applicable to other antigens as well.
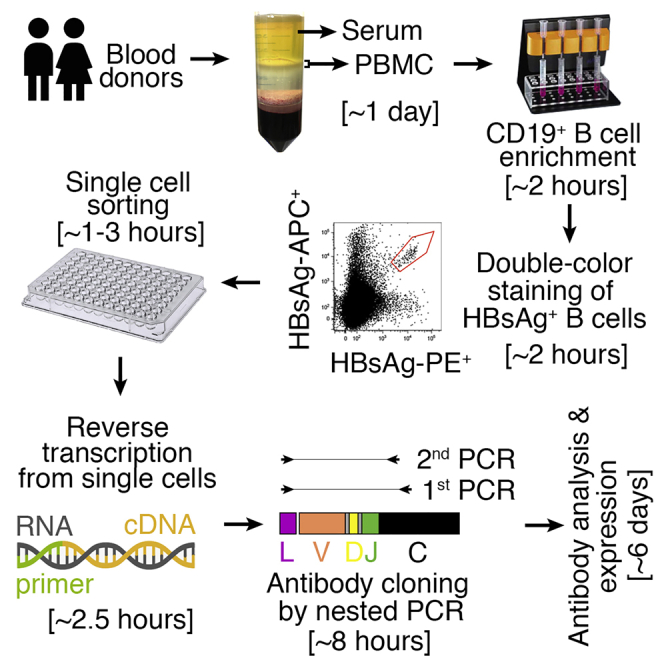
Figure 5. Diagram of Single-Cell Sorting and Cloning of HBsAg-Binding Memory B Cells from Human Peripheral Blood Mononuclear Cells [7]
4. Guinea Pig Antigen-Specific Single B Cell Sorting and Monoclonal Antibody Cloning
A study established a platform for sorting antigen-specific single B-cells and cloning mAbs, aimed at analyzing immunization- or viral infection-elicited antibody response at the clonal level in guinea pigs. Researchers stained PBMCs from a guinea pig immunized with the HIV-1 envelope glycoprotein trimer analog (BG505 SOSIP), using anti-guinea pig IgG and IgM fluorochrome conjugates, along with fluorochrome-conjugates BG505 SOSIP trimer as antigen (Ag) probes to sort for Ag-specific IgGhi IgMlo B cells at single cell density. The team designed a set of guinea pig Ig gene-specific primers to amplify cDNAs encoding B cell receptor variable regions [V(D)J segments] from the sorted Ag-specific B cells. After validating these sequences by sequencing and annotating them using IgBLAST, they were cloned into expression vectors with human IgG1 constant regions for Ig heavy and light chains, and co-transfected into 293F cells to reconstruct full-length antibodies in guinea pig-human chimeric IgG1 form. From 88 isolated antigen-specific B-cells, 24 (27%) with native-paired heavy and light chains were recovered. Additionally, 85% of the recombinant mAbs demonstrated positive binding to antigen probes in enzyme-linked immunosorbent assays and/or BioLayer interferometry assays, with five antibodies from four clonal lineages neutralizing the HIV-1 tier 1 virus ZM109. Overall, this technique combining Ag-specific single B cell sorting with gene-specific single-cell RT-PCR proved highly efficient and accurate, facilitating future mAb isolation and analysis of B cell responses to infection or immunization in guinea pig models.
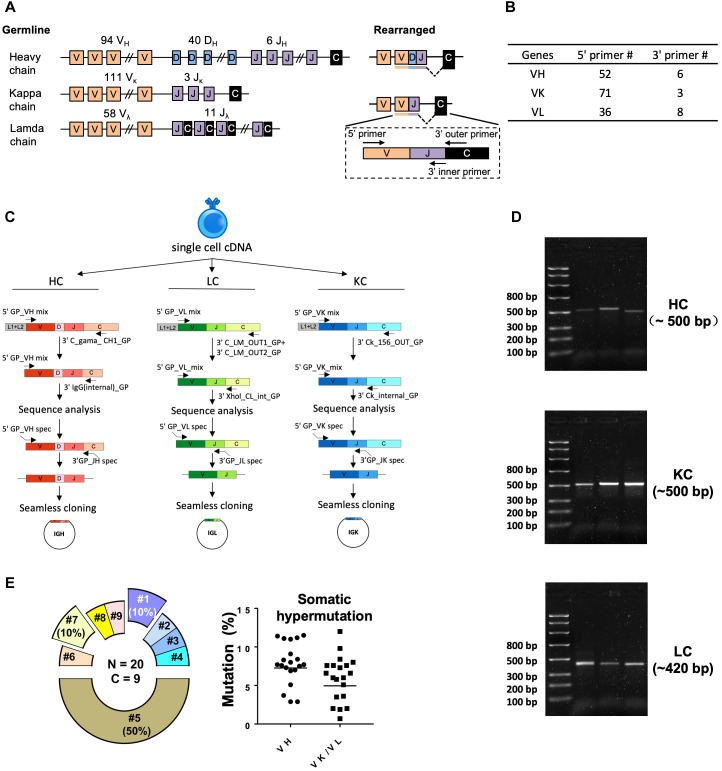
Figure 6. Single-Cell RT-PCR Amplification of Antigen-Specific Guinea Pig B-Cell IGH, IGL, and IGK Transcripts[8]
Sample Submission Requirements
1. Protein Purity > 90%
2. Minimization of Impurity Contamination
Services at MtoZ Biolabs
1. Comprehensive Experimental Procedures
2. Relevant Instrumentation Parameters
3. Raw Experimental Data
4. Data Analysis Report
Applications
1. Single B-Cell Antibody Library Research
Analysis based on classical mouse and human antibody libraries, which is conducted at the cellular level, has markedly advanced our understanding of universal B-cell immunology principles. Initially, the study of Ig genes from individual B cells via single-cell PCR at the genomic level was aimed at clarifying the molecular mechanisms underlying Ig gene rearrangement and allelic exclusion events. This system has been further developed to isolate valuable mAbs from immunized donors for potential therapeutic applications, epitope mapping, or vaccine design.
FAQ
Q1: What are the common gating strategies for different types of B cells?
Differentiation is based on the expression of surface markers on B cells. Naive B cells (CD19+ CD3- CD27lo CD38int), memory B cells (CD19+ CD3- CD27lo CD38int), and plasmablasts (CD19+ CD3- CD27lo CD38int).
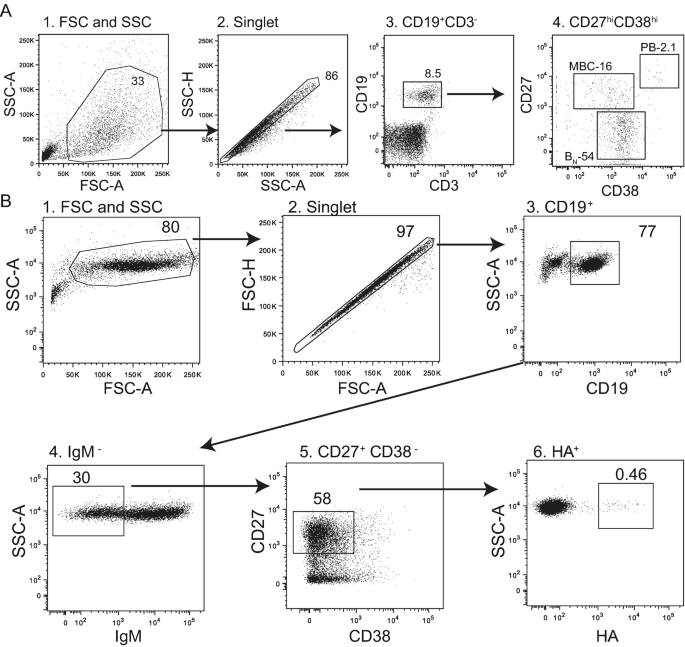
Figure 7. Strategy for B-Cell Gating[9]
References
[1] Snapkov I, Chernigovskaya M, Sinitcyn P, Lê Quý K, Nyman TA, Greiff V. Progress and challenges in mass spectrometry-based analysis of antibody repertoires. Trends Biotechnol. 2022 Apr;40(4):463-481. doi: 10.1016/j.tibtech.2021.08.006. Epub 2021 Sep 14. PMID: 34535228.
[2] de Brito PM, Saruga A, Cardoso M, Goncalves J. Methods and cell-based strategies to produce antibody libraries: current state. Appl Microbiol Biotechnol. 2021 Oct;105(19):7215-7224. doi: 10.1007/s00253-021-11570-x. Epub 2021 Sep 15. PMID: 34524471.
[3] Alejandra WP, Miriam Irene JP, Fabio Antonio GS, Patricia RR, Elizabeth TA, Aleman-Aguilar JP, Rebeca GV. Production of monoclonal antibodies for therapeutic purposes: A review. Int Immunopharmacol. 2023 Jul;120:110376. doi: 10.1016/j.intimp.2023.110376. Epub 2023 May 25. PMID: 37244118.
[4] von Boehmer L, Liu C, Ackerman S, Gitlin AD, Wang Q, Gazumyan A, Nussenzweig MC. Sequencing and cloning of antigen-specific antibodies from mouse memory B cells. Nat Protoc. 2016 Oct;11(10):1908-1923. doi: 10.1038/nprot.2016.102. Epub 2016 Sep 15. PMID: 27658009.
[5] Starkie DO, Compson JE, Rapecki S, Lightwood DJ. Generation of Recombinant Monoclonal Antibodies from Immunised Mice and Rabbits via Flow Cytometry and Sorting of Antigen-Specific IgG+ Memory B Cells. PLoS One. 2016 Mar 29;11(3):e0152282. doi: 10.1371/journal.pone.0152282. PMID: 27022949; PMCID: PMC4811437.
[6] Goldstein, L.D., Chen, YJ.J., Wu, J. et al. Massively parallel single-cell B-cell receptor sequencing enables rapid discovery of diverse antigen-reactive antibodies. Commun Biol 2, 304 (2019). https://doi.org/10.1038/s42003-019-0551-y.
[7] Zhou Y, Liu Z, Wang Z, Zhang Q, Mayer CT, Schoofs T, Nussenzweig MC, de Jong YP, Wang Q. Single-Cell Sorting of HBsAg-Binding Memory B Cells from Human Peripheral Blood Mononuclear Cells and Antibody Cloning. STAR Protoc. 2020 Oct 16;1(3):100129. doi: 10.1016/j.xpro.2020.100129. PMID: 33377023; PMCID: PMC7757113.
[8] Lei L, Tran K, Wang Y, Steinhardt JJ, Xiao Y, Chiang CI, Wyatt RT, Li Y. Antigen-Specific Single B Cell Sorting and Monoclonal Antibody Cloning in Guinea Pigs. Front Microbiol. 2019 Apr 23;10:672. doi: 10.3389/fmicb.2019.00672. PMID: 31065249; PMCID: PMC6489837.
[9] Guthmiller JJ, Dugan HL, Neu KE, Lan LY, Wilson PC. An Efficient Method to Generate Monoclonal Antibodies from Human B Cells. Methods Mol Biol. 2019;1904:109-145. doi: 10.1007/978-1-4939-8958-4_5. PMID: 30539468.
MtoZ Biolabs, an integrated chromatography and mass spectrometry (MS) services provider.
Related Services
How to order?







