SILAC vs. TMT: Comparative Analysis in Quantitative Subcellular Proteomics
-
SILAC- and TMT-based quantitative subcellular proteomics services.
-
Customized experimental design and sample preparation guidance.
-
Integrated analysis of subcellular localization data with GO/KEGG pathway enrichment.
-
AI-assisted interpretation of subcellular protein distribution patterns.
Cells are highly compartmentalized systems in which distinct subcellular organelles, such as the nucleus, mitochondria, endoplasmic reticulum, and Golgi apparatus, perform specialized biological functions. Subcellular proteomics focuses on elucidating the spatial distribution and dynamic behavior of proteins within the cellular environment and plays a critical role in advancing our understanding of signal transduction, metabolic reprogramming, disease pathogenesis, and drug target discovery. Achieving high-resolution subcellular localization analysis requires not only efficient cell fractionation strategies but also accurate and robust quantitative proteomics approaches. Currently, stable isotope labeling by amino acids in cell culture (SILAC) and tandem mass tag (TMT) labeling represent the two most widely adopted strategies for quantitative subcellular proteomics.
Basic Principles of SILAC and TMT
1. SILAC: A Metabolic Labeling Strategy
SILAC (Stable Isotope Labeling by Amino acids in Cell culture) introduces light or heavy isotope-labeled amino acids (e.g., ^13C₆-Lys and ^13C₆-Arg) during cell culture, enabling the in vivo synthesis of isotopically labeled proteins. This strategy is particularly suitable for quantitative analyses in mammalian cell lines. Because isotopic labels are incorporated at the protein synthesis stage, SILAC offers near-complete labeling efficiency and highly linear quantitative performance.
(1) Advantages: Endogenous metabolic labeling minimizes technical variability between samples and is well suited for experiments involving dynamic cellular processes.
(2) Limitations: SILAC is not applicable to primary cells, tissue specimens, or cell types that are difficult to culture.
2. TMT: A Chemical Labeling Strategy
TMT (Tandem Mass Tag) is a chemical labeling approach in which isobaric tags are covalently attached to peptide N-termini or lysine side chains, allowing multiplexed quantification of different samples. Modern TMT reagents typically support 10-18 labeling channels, providing substantial advantages for large-scale and high-throughput comparative analyses.
(1) Advantages: High multiplexing capacity makes TMT particularly suitable for large-scale clinical and cohort-based studies.
(2) Limitations: Chemical labeling may introduce batch effects, and quantitative accuracy can be affected by ratio compression.
Differences in Quantitative Accuracy and Spatial Resolution
Subcellular proteomics demands precise quantification of low-abundance proteins whose localization is highly sensitive to cellular perturbations. A comparison of SILAC and TMT across key performance metrics is summarized below:
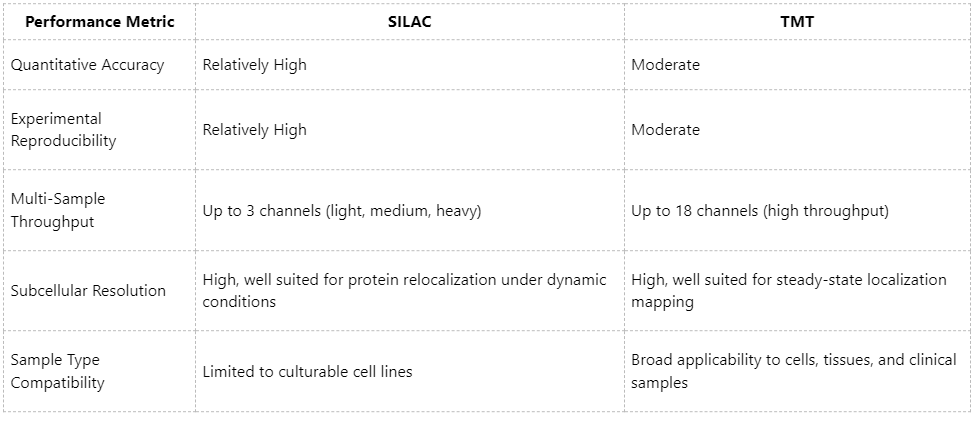
Because SILAC does not require additional processing steps before or after labeling, it more faithfully reflects endogenous protein distributions and therefore performs particularly well in studies of dynamic subcellular relocalization, such as stimulus responses or cell cycle-dependent changes. In contrast, TMT is more advantageous for constructing steady-state subcellular localization atlases, especially in large-scale parallel analyses.
Comparison of Application Scenarios: Best Practices for Different Strategies
1. Applications Best Suited for SILAC
(1) Studies of stimulus-induced protein translocation, including mitochondrial or nuclear protein redistribution following cellular stress or pharmacological treatment.
(2) Analyses of protein-protein interaction-associated localization changes, such as the assembly and disassembly of protein complexes under varying conditions.
2. Applications Best Suited for TMT
(1) Construction of subcellular protein localization landscapes in tissue samples, such as comprehensive mitochondrial proteome maps across different cancer types.
(2) Spatial proteomic profiling under high-throughput drug treatment conditions, enabling simultaneous comparison of multiple experimental perturbations.
Trends in Technical Integration: SILAC × TMT × AI Spatial Proteomics
Driven by advances in deep learning-based subcellular localization prediction tools (e.g., DeepLoc and SubCell-BERT) and high-resolution mass spectrometry, quantitative subcellular proteomics is increasingly moving toward multidimensional data integration. Emerging studies have explored the combined use of SILAC and TMT to integrate the quantitative precision of endogenous metabolic labeling with the scalability of multiplexed analysis. When coupled with AI-based clustering of protein localization patterns, these approaches facilitate systematic investigations of the spatial organization of disease-associated biomarkers.
Service Advantages of MtoZ Biolabs in Subcellular Proteomics
At MtoZ Biolabs, we integrate multiple mass spectrometry platforms (Orbitrap Exploris, Fusion Lumos, and timsTOF Pro) with optimized subcellular fractionation workflows to provide:
Whether the research focus is spatial reprogramming of signaling pathways or subcellular proteome remodeling under disease conditions, MtoZ Biolabs delivers high-quality, publication-ready datasets.
SILAC and TMT offer complementary strengths in subcellular proteomics and should be selected based on specific experimental objectives rather than viewed as interchangeable approaches. A clear understanding of their underlying principles and application contexts enables the rational design of experiments and enhances the spatial interpretability of proteomic data. Researchers are encouraged to collaborate with MtoZ Biolabs to tailor subcellular proteomics solutions that best meet their project requirements.
MtoZ Biolabs, an integrated chromatography and mass spectrometry (MS) services provider.
Related Services
How to order?







