RNA Pull-Down Service
- Identification of RNA-binding proteins for novel long noncoding RNAs and circular RNAs
- Functional characterization of untranslated regions and regulatory RNA elements
- Validation of predicted RNA protein interactions from computational analyses
- Investigation of RNA-mediated post-transcriptional regulation
- Mapping of viral RNA host protein interactomes
- Mechanistic studies of disease-associated RNA variants
- Support for RNA-targeted drug discovery and target validation
- Cell-Based Inputs: cultured cells, including adherent or suspension formats, as well as virus-infected cells when relevant
- Tissue-Derived Inputs: tissue homogenates or tissue lysates
- Subcellular Fractions: cytosolic, nuclear, or other enriched fractions to improve specificity for compartmentalized RNAs or RBPs
- Cell-Free Systems: reconstituted RNP mixtures or purified components
MtoZ Biolabs provides a comprehensive RNA Pull-Down Service designed to identify and characterize RNA-binding proteins and RNA-associated protein complexes with high specificity and confidence. This service supports studies of RNA protein interactions, post-transcriptional regulation, RNA-mediated signaling, and disease-relevant RNA regulatory mechanisms across diverse biological systems.
What Is RNA Pull-Down?
Messenger RNAs, long noncoding RNAs, circular RNAs, and regulatory RNA elements exert their biological roles primarily through interactions with proteins. These RNA protein interactions govern RNA stability, localization, translation efficiency, modification, and turnover, thereby shaping cellular phenotypes and responses.
RNA pull-down analysis is a targeted biochemical strategy used to capture proteins that directly or indirectly associate with a defined RNA sequence. By enriching RNA-bound protein complexes from biological samples and identifying them using mass spectrometry or immunodetection, RNA pull-down enables mechanistic investigation of RNA-centric regulatory networks.
This approach is particularly valuable for decoding the functional partners of novel noncoding RNAs, mapping RNA regulatory hubs, validating predicted RNA protein interactions, and uncovering disease-associated RNA interactomes.
RNA Pull-Down Service at MtoZ Biolabs
MtoZ Biolabs provides two complementary RNA pull-down strategies that address different biological questions and experimental constraints. These approaches are optimized to balance specificity, physiological relevance, and analytical depth.
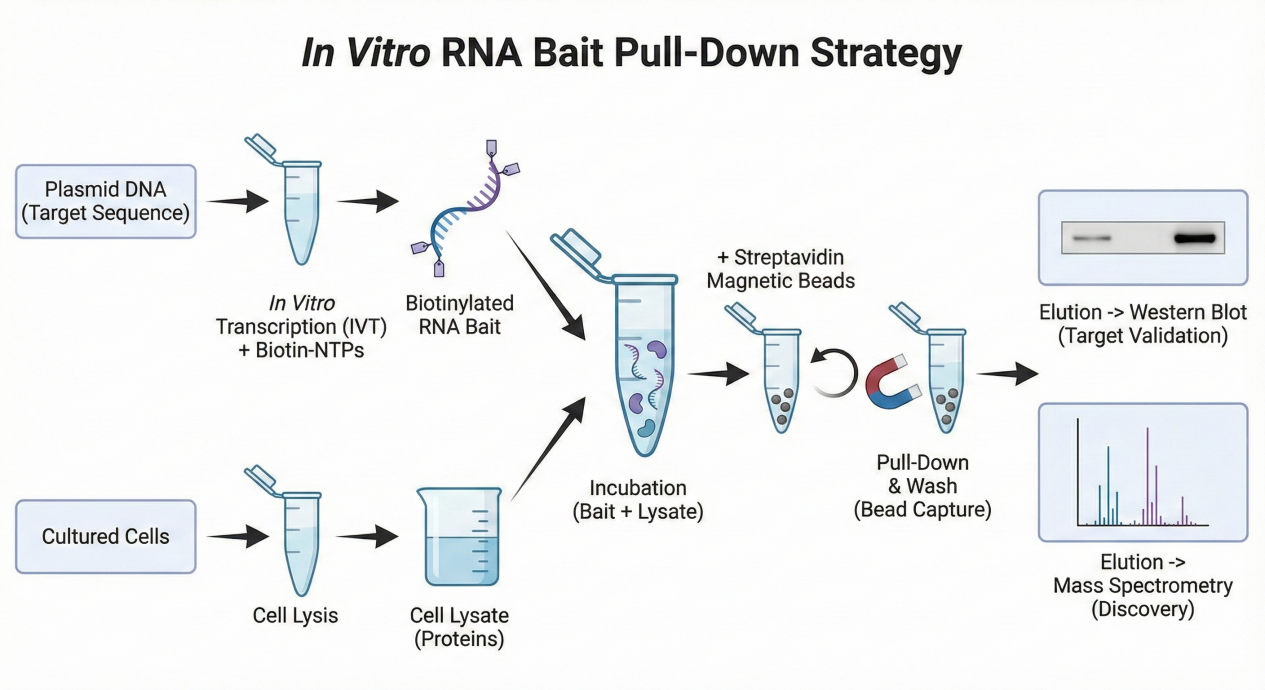
1. In Vitro RNA Bait Pull-Down Strategy
The in vitro RNA bait pull-down approach uses synthetically prepared or in vitro transcribed RNA molecules as affinity probes. The RNA is typically labeled or conjugated to an affinity handle, allowing it to be immobilized on beads and incubated with protein extracts.
Key features of this strategy include:
🔸Precise control over RNA sequence length and composition
🔸Ability to interrogate specific RNA motifs, domains, or mutations
🔸Flexible experimental conditions for binding optimization
🔸Compatibility with diverse sample sources, including nuclear, cytoplasmic, or tissue lysates
This method is particularly suitable for dissecting direct RNA protein interactions, comparing wild-type and mutant RNA variants, and screening candidate RNA-binding proteins under defined conditions.
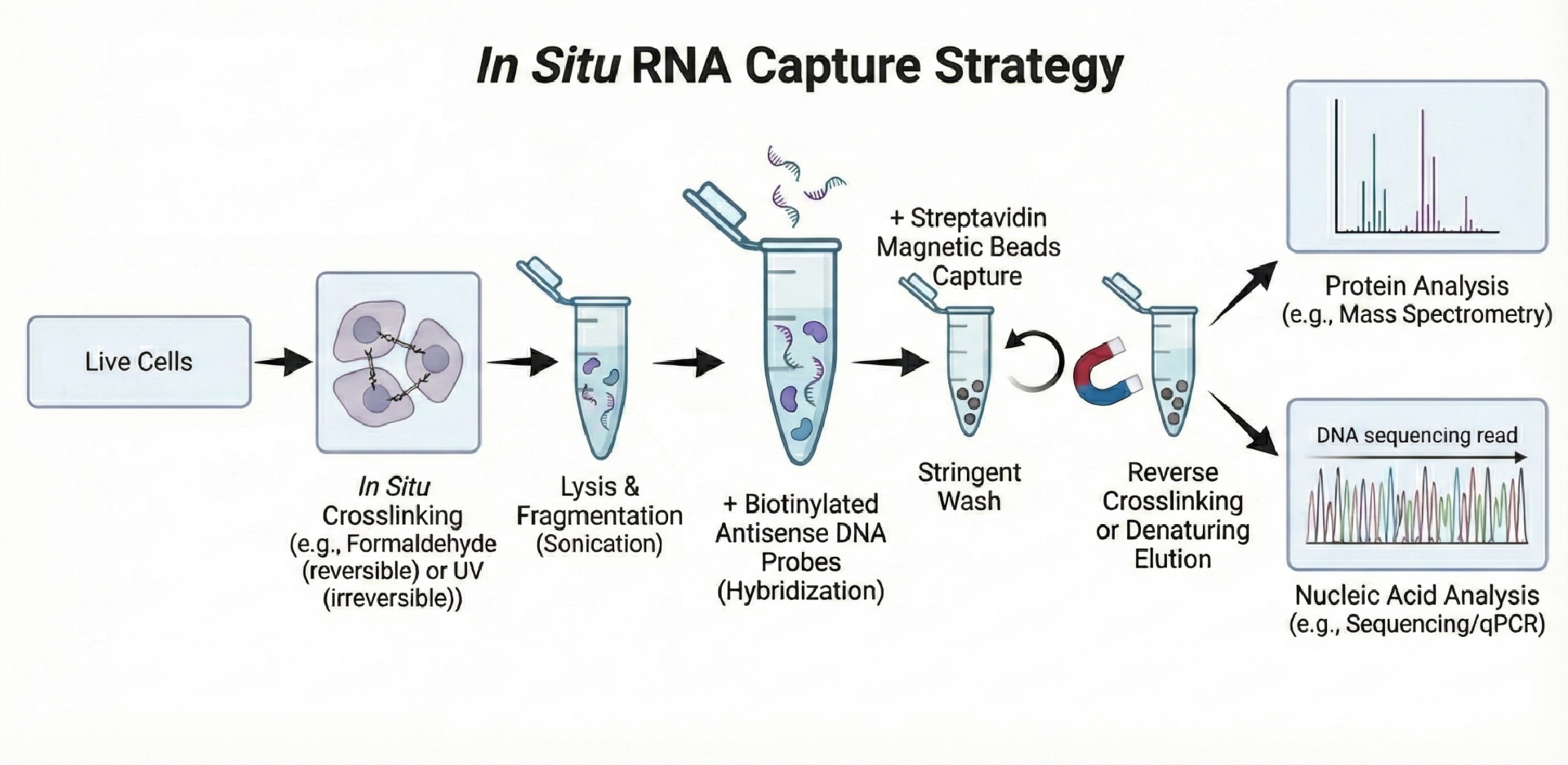
2. In Situ RNA Capture Strategy
The in situ RNA capture strategy preserves RNA protein interactions formed in the native cellular environment prior to isolation. RNA protein complexes are stabilized within intact cells, followed by selective capture of the target RNA and its associated proteins.
Distinct advantages of this approach include:
🔸Retention of physiological RNA protein interactions
🔸Reduced disruption of transient or weakly associated complexes
🔸Improved relevance for functional and disease-related studies
🔸Applicability to endogenous RNAs without extensive manipulation
This strategy is well suited for studying dynamic RNA interactomes, condition-dependent binding events, and context-specific regulatory complexes that may be lost in in vitro systems.
Workflow of RNA Pull-Down Service
The RNA Pull-Down Service at MtoZ Biolabs follows a streamlined and reproducible analytical workflow designed to maximize interaction specificity while minimizing background noise.
1. Experimental design and RNA strategy selection based on research objectives
2. RNA preparation or in situ capture optimization
3. Controlled incubation of RNA bait with biological samples to allow specific RNA protein interactions
4. Affinity enrichment of RNA-associated protein complexes
5. Stringent washing to reduce nonspecific binding
6. Protein elution and digestion
7. High-resolution LC-MS MS analysis for protein identification
8. Data processing, filtering, and biological interpretation
Why Choose MtoZ Biolabs
✔️Dual RNA Pull-Down Strategies
Supports both in vitro RNA bait pull-down and in situ RNA capture to match different biological contexts and research goals.
✔️Interaction-Focused Optimization
Binding, incubation, and enrichment conditions are optimized to preserve specific RNA protein interactions while minimizing background.
✔️Direct Integration with LC-MS MS
Affinity-enriched RNA protein complexes are analyzed using high-resolution LC-MS MS for unbiased protein identification.
✔️Strict Background Control
Experimental controls and data filtering are applied to distinguish true RNA interactors from nonspecific binders.
Applications of RNA Pull-Down Service
RNA Pull-Down Service at MtoZ Biolabs supports a wide range of research applications across basic biology, translational research, and therapeutic development. Representative applications include:
FAQ
Q1: What types of samples are suitable?
If you have special or limited material, we can assess feasibility and optimize a suitable workflow.
Q2: How should I prepare my samples?
1. Handling and Consistency: maintain consistent handling across groups and keep samples cold during processing
2. Lysis and Buffer Expectations: use RNase-free lysis buffers, avoid harsh denaturants, and include RNase inhibitors; keep glycerol low if used
3. Additives and Interaction Stability: Mg2+ can be used when structural preservation is important
4. Storage and Shipping: snap-freeze lysates promptly, store at low temperature, ship on dry ice, and avoid repeated freeze-thaw cycles
5. Information to Include: provide buffer composition, basic RNA integrity notes if available, and your research objective to guide controls and enrichment strategy
For more information, please refer to Sample Submission Guidelines for Proteomics and Sample Submission Guidelines for Metabolomics.
Q3: What is the service general workflow?
Q4: What data formats are provided?
1. Raw Data Files: vendor native MS files and mzML
2. Processed Tables: XLSX or CSV (protein IDs and gene symbols, peptide evidence, quantitative metrics, confidence values, enrichment and statistics versus controls when relevant)
3. Figures: PDF plus PNG or TIFF (QC summaries, replicate agreement, enrichment plots, heatmaps, and network diagrams)
4. Annotations: XLSX or CSV (pathways, complexes, domain or RNA-binding feature notes, and background or contaminant flags)
5. Optional Add-Ons: targeted confirmation outputs in XLSX or CSV with supporting PDF and PNG or TIFF figures when included
6. Comprehensive Result Reports: PDF (key protocol settings, capture and wash conditions, elution approach, instrument settings, software versions, results, and interpretation)
Additional formats can be provided upon request to meet specific analysis or publication requirements.
Start Your Project with MtoZ Biolabs
Whether you are exploring the functional partners of a newly discovered RNA or validating RNA-mediated regulatory mechanisms, MtoZ Biolabs RNA Pull-Down Service provides a robust and flexible platform for RNA protein interaction analysis. Our expertise, analytical rigor, and collaborative approach help transform RNA interaction hypotheses into actionable biological insights.
Contact us to discuss your experimental design or request a quote. Our technical specialists are available to provide a free business assessment.
How to order?







