RNA Proximity Labeling Service
MtoZ Biolabs provides an RNA Proximity Labeling Service that maps the in situ protein microenvironment surrounding a target RNA in living cells, enabling sensitive identification of proximal RNA-associated proteins by proximity labeling and LC-MS/MS. The workflow is well suited for capturing transient, low-affinity, or spatially confined interactions and supports studies of diverse endogenous and exogenous RNAs across mechanistic biology, functional genomics, and drug discovery.
What Is RNA Proximity Labeling?
Most RNAs operate within dynamic ribonucleoprotein assemblies that change with localization, cell state, stress, and treatment. Classic protein-RNA interaction analysis strategies such as RNA pull-down, RIP, or CLIP are powerful, but they can be limited by disruption of weak interactions during purification, dependence on crosslinking conditions, and challenges in resolving context-specific or compartment-restricted interactions.
RNA proximity labeling addresses these gaps by tagging proteins near an RNA of interest directly in cells, then using affinity enrichment and mass spectrometry to identify labeled proteins. This approach is particularly valuable when you want to capture short-lived interactions, distinguish local neighborhoods, or interrogate RNA-associated proteomes under physiological perturbations.
Principle of RNA Proximity Labeling
RNA proximity labeling is an in vivo labeling strategy that couples an RNA-targeting module with an enzyme that generates reactive labeling species. When the targeting module brings the enzyme close to the RNA of interest, nearby proteins are covalently tagged, most commonly with biotin or peroxidase. Biotinylated proteins are then enriched using streptavidin beads and identified by LC-MS/MS.
Key components of the method include:
1. RNA targeting to position the labeling enzyme near the RNA of interest
2. Enzymatic labeling to covalently tag proteins within a short effective radius
3. Streptavidin enrichment to isolate labeled proteins under stringent conditions
4. Mass spectrometry to identify and quantify RNA-proximal proteins
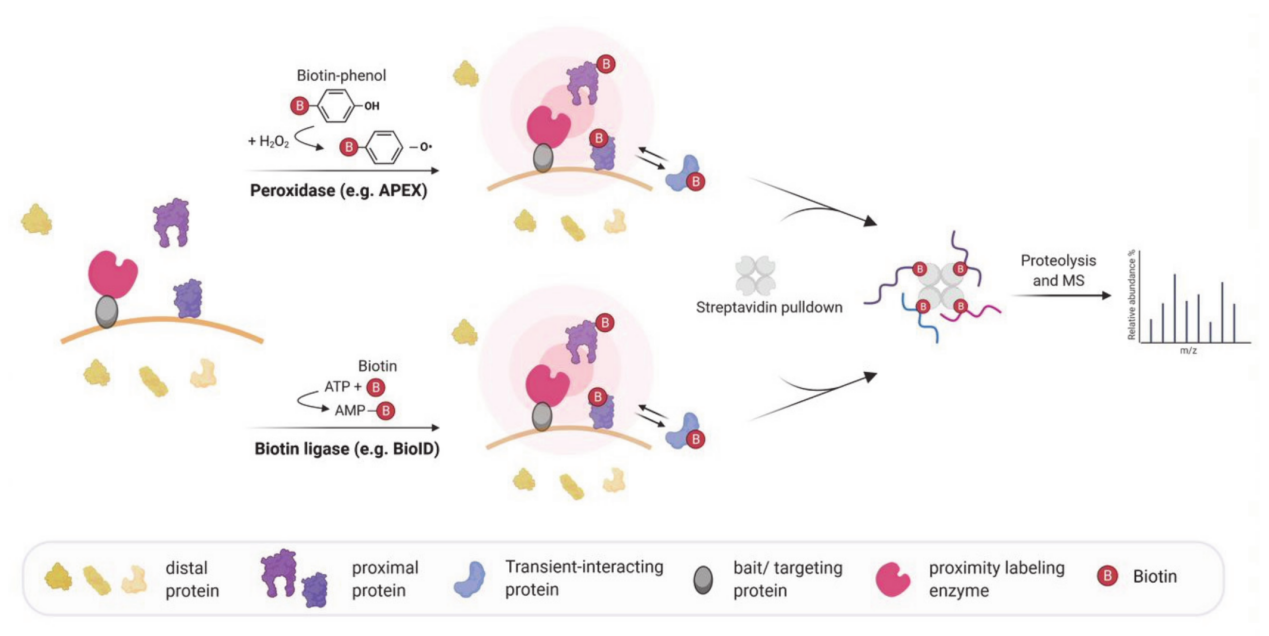
Weissinger, R. et al. Molecules. 2021.
Figure 1. Schematic Workflow of Peroxidase and Biotin Ligase Based In Vivo Proximity Labeling for Mapping Molecular Interactions
RNA Proximity Labeling Service at MtoZ Biolabs
MtoZ Biolabs delivers an integrated workflow for RNA proximity labeling, from targeting strategy planning through proteomic identification and data interpretation. We help select appropriate targeting and labeling configurations, optimize labeling conditions and controls, perform streptavidin enrichment, and apply LC-MS/MS-based discovery to generate a prioritized set of RNA-proximal protein candidates.
Service capabilities include:
🔸Baseline profiling of the RNA-proximal proteome under a defined condition
🔸Comparative profiling across treatment, stress, differentiation, genotype, or time course designs
🔸Quantitative analysis options to support multi-group studies and candidate ranking
🔸Bioinformatic filtering and enrichment scoring to improve confidence and interpretability
Why Choose MtoZ Biolabs
✔️Expert Scientific Team
Projects are supported by experienced scientists with deep expertise in RNA biology, proteomics, and genomics.
✔️Advanced Platforms
Robust experimental workflows and high-performance analytical platforms enable sensitive and reproducible results.
✔️Customizable Service Options
Flexible study designs and modular deliverables tailored to your RNA targets, sample types, and research objectives.
✔️Fast Turnaround
Streamlined execution and clear project management to keep timelines efficient without compromising quality.
✔️Outstanding Customer Support
Responsive communication, technical guidance, and reliable reporting throughout the project lifecycle.
Applications of RNA Proximity Labeling Service
1. lncRNA and CircRNA Mechanism Studies
Identify protein partners and local protein environments that support chromatin regulation, nuclear body formation, or RNA-mediated scaffolding.
2. MRNA Localization and Translation Regulation
Define proteins associated with localized mRNAs, stress granule related environments, or translation initiation contexts.
3. Viral RNA Host Factor Discovery
Map host proteins proximal to viral RNAs during infection models to support host factor identification and antiviral mechanism studies.
4. Drug Discovery and Target Validation
Evaluate how perturbations alter RNA-proximal protein neighborhoods, supporting mechanism of action hypotheses for RNA-targeting therapeutics or pathway modulators.
5. Condition-Dependent Interactome Remodeling
Compare RNA-proximal proteomes across stress, differentiation, treatment, or genotype to identify context-specific regulators.
FAQ
Q1: What types of samples are suitable?
1. Cells: cultured cells from defined conditions, including treatment groups or engineered lines
2. Tissues: fresh or properly frozen tissues
3. Purified RNA: purified target RNA or total RNA
For other sample types or low-yield materials, please contact MtoZ Biolabs in advance for customized preparation guidance.
Q2: How should I prepare my samples?
1. Handling and processing: Collect samples as consistently as possible across groups and minimize time at room temperature. Use RNase-free consumables and keep workflows cold to protect RNA integrity.
2. Aliquoting and labeling: Aliquot to avoid repeated freeze-thaw cycles. Label each tube clearly with sample ID, group, replicate, and sample type, and include a matching sample sheet.
3. Storage: Store samples at -80°C whenever applicable and keep them frozen until shipment. Avoid unnecessary temperature fluctuations during storage and transfer.
4. Shipping: Ship samples on dry ice. Ensure sufficient dry ice for the entire transit window.
For more information, please refer to Sample Submission Guidelines for Proteomics and Sample Submission Guidelines for Metabolomics.
Q3: What is the service general workflow?
Q4: What data formats are provided?
1. Raw data: vendor MS raw files plus optional mzML exports
2. Search and identification outputs: standard database search result files as applicable
3. Processed tables: XLSX or CSV for protein IDs, peptide evidence, quantitative metrics, enrichment versus controls, and statistics when comparative designs are included
4. Figures and QC: PDF plus PNG or TIFF (QC summaries, replicate correlation, enrichment plots, heatmaps, network views when applicable)
5. Report and methods: concise PDF describing assay design, controls, key parameters, instrument settings, and software versions for reproducibility
Additional formats can be provided upon request to meet specific analysis or publication requirements.
Start Your Project with MtoZ Biolabs
Contact us to discuss your experimental design or request a quote. Our technical specialists are available to provide a free business assessment.
How to order?







