Quantitative Phosphoproteomics: Comparing SILAC, TMT, and Label-Free Methods
- Endogenous labeling ensures high quantitative accuracy and is well-suited for detecting subtle phosphorylation changes.
- When combined with phosphopeptide enrichment, SILAC exhibits excellent reproducibility.
- The workflow is straightforward and avoids biases associated with variable labeling efficiency.
- Applicable only to cell lines that can be cultured in vitro.
- Limited sample throughput, typically restricted to comparisons of 2–3 conditions.
- Not applicable to tissue-derived or clinical samples.
- High multiplexing capacity, enabling simultaneous analysis of 6–18 samples.
- Broad applicability to diverse sample types, including tissues, cultured cells, and clinical specimens.
- Well-established data analysis pipelines and mature commercial service support.
- Particularly suitable for quantitative phosphoproteomics, as enriched samples analyzed in a single run can effectively reduce batch-related variability.
- Relatively high reagent and experimental costs.
- Susceptibility to ratio compression effects, which may underestimate true fold changes.
- Requires accurate sample quantification and carefully optimized mass spectrometry acquisition strategies.
- No labeling is required, providing high flexibility and suitability for complex or limited sample types.
- Scalable to comparative analyses involving dozens of samples.
- Lower overall cost relative to labeling-based approaches.
- Separate analysis of samples introduces pronounced batch effects.
- Strong dependence on instrument stability and robust data normalization algorithms.
- Limited suitability for reliable detection and quantification of ultra-low-abundance phosphopeptides.
In phosphoproteomics research, identification of phosphorylation sites alone is insufficient. Quantitative information is the primary driver of biological insight, addressing questions such as which sites undergo significant changes upon stimulation and which signaling pathways are most actively regulated by phosphorylation. These analyses rely critically on accurate and reproducible quantitative strategies. Quantitative phosphoproteomics not only facilitates the discovery of potential biomarkers but also supports the identification of therapeutic targets, thereby contributing to the advancement of precision medicine. In addition, this approach is widely applied in studies of drug mechanisms of action and therapeutic efficacy assessment, making it a key analytical tool in modern drug development. With continuous advances in mass spectrometry instrumentation and phosphopeptide enrichment techniques, the sensitivity and coverage of quantitative phosphoproteomics have markedly improved, enabling analyses with increased depth and scope. Consequently, quantitative phosphoproteomics has become an indispensable component of systems biology research and plays a central role in elucidating complex biological processes and disease-associated regulatory features. Currently, three major quantitative strategies are widely employed: SILAC (stable isotope labeling), TMT (isobaric tag-based quantification), and label-free quantification (LFQ).
Overview of Quantitative Phosphoproteomics Methods
1. SILAC: Metabolic Labeling at the Cell Culture Level
SILAC (Stable Isotope Labeling by Amino acids in Cell culture) is a metabolic labeling strategy in which light or heavy isotope-labeled amino acids (e.g., Lys-0 versus Lys-8) are supplemented into cell culture media. During protein biosynthesis, cells endogenously incorporate these labeled amino acids into newly synthesized proteins, enabling accurate and internally controlled quantitative comparisons.
(1) Advantages
(2) Limitations
2. TMT: Multiplex Chemical Labeling for Enhanced Throughput
TMT (Tandem Mass Tag) is a chemical labeling approach in which isobaric tags are introduced at the peptide level after enzymatic digestion. Individually labeled samples are subsequently combined and analyzed in a single mass spectrometry run, with quantitative information derived from reporter ions released during MS/MS fragmentation.
(1) Advantages
(2) Limitations
3. Label-Free Quantification: Flexible but Highly Instrument-Dependent
Label-free quantification involves independent mass spectrometry analysis of individual samples, followed by quantitative comparison based on peptide signal intensities or spectral counting metrics. By eliminating labeling steps, this strategy offers a streamlined workflow and is particularly advantageous for large-scale or exploratory studies.
(1) Advantages
(2) Limitations
Key Comparative Metrics for Quantitative Phosphoproteomics
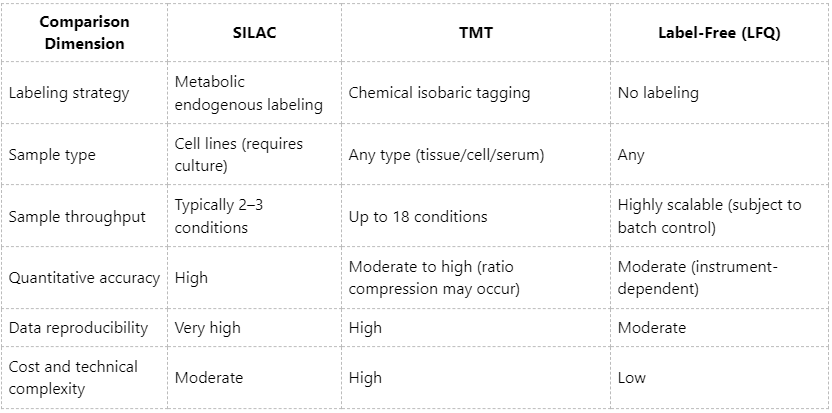
Selecting an Appropriate Quantitative Strategy for Phosphoproteomics Studies
1. Selection Based on Research Objectives
(1) Mechanistic studies and signaling pathway analysis: SILAC or TMT is recommended. SILAC is particularly suited for precise quantification in cell-based systems, whereas TMT supports broader comparative analyses across multiple sample types.
(2) Differential analysis of clinical samples and biomarker discovery: TMT or LFQ is generally preferred to accommodate tissue-derived samples and high-throughput study designs.
(3) High-sensitivity phosphosite profiling: TMT combined with tandem phosphopeptide enrichment strategies (e.g., IMAC followed by TiO₂) currently represents a widely adopted solution.
2. Considerations of Budget and Sample Accessibility
(1) Adequate budget with a requirement for high throughput: TMT is the preferred option.
(2) Limited budget or challenges in unified sample processing: Label-free quantification, combined with downstream bioinformatics optimization, can be considered.
(3) Well-controlled cell line–based experimental models: SILAC remains one of the most accurate quantitative strategies available.
In phosphoproteomics research, the selection of a quantitative phosphoproteomics strategy not only influences data quality but also directly impacts experimental design and the depth of biological interpretation. If guidance on quantitative strategy selection is required during project planning, consultation with MtoZ Biolabs is welcome. We will provide an integrated phosphoproteomics quantification solution tailored to your sample types, research objectives, and budget.
MtoZ Biolabs, an integrated chromatography and mass spectrometry (MS) services provider.
Related Services
How to order?







