Principles of Phosphorylated Protein Analysis: Unlocking Cellular Signaling Pathways
The precise transmission of intracellular and extracellular signals depends on a complex network of regulatory mechanisms. Protein phosphorylation, one of the most critical post-translational modifications, functions as a central molecular regulatory language that governs core biological processes ranging from metabolic regulation to cell division, apoptosis, and migration. Protein phosphorylation is catalyzed by protein kinases, which covalently attach a phosphate group (PO₄³⁻) to specific amino acid residues, predominantly serine (Ser), threonine (Thr), or tyrosine (Tyr). Protein phosphatases remove these phosphate groups, forming a highly reversible and tightly controlled regulatory system. In studies of cancer, immunity, autoimmune disorders, and neurodegenerative diseases, researchers increasingly rely on phosphorylated protein analysis to reconstruct the dynamic architecture of cellular signaling pathways. Owing to its rapid and precise regulatory properties, phosphorylation plays a central role in signal transduction networks.
Analytical Principles: How to Identify Phosphorylation Signals Within Complex Peptide Mixtures?
The primary objective of phosphorylated protein analysis is to identify and quantify phosphorylation sites from complex biological samples and map them onto regulatory networks through bioinformatic approaches. The overall workflow can be divided into five major steps:
1. Protein Extraction and Enzymatic Digestion
Total proteins are rapidly extracted using lysis buffers containing phosphatase inhibitors, followed by enzymatic digestion with trypsin to generate short peptides suitable for mass spectrometry analysis.
2. Phosphopeptide Enrichment
Because phosphopeptides typically account for less than 2% of total peptides, enrichment is required prior to mass spectrometric detection.
Common strategies include:
(1) TiO₂ enrichment: capture based on electrostatic interactions between metal oxides and phosphate groups
(2) IMAC (Immobilized Metal Affinity Chromatography)
(3) pTyr antibody affinity purification: specifically targeting tyrosine phosphorylation sites, offering high specificity but at higher cost
(4) Sequential enrichment (SIMAC): fractionation according to the degree of phosphorylation to improve identification efficiency of low-abundance multiphosphorylated peptides
3. LC-MS/MS Analysis
Enriched phosphopeptides are separated by liquid chromatography and subsequently analyzed using high-resolution mass spectrometry (such as Orbitrap instruments). During MS/MS fragmentation, mass-to-charge ratios (m/z) and retention times are recorded, enabling identification of peptide sequences and precise localization of phosphorylation sites.
DDA vs. DIA Modes
(1) DDA (Data-Dependent Acquisition): particularly suited for exploratory studies and spectral library construction
(2) DIA (Data-Independent Acquisition): well suited for multi-sample quantitative analysis, offering enhanced reproducibility
4. Data Processing and Site Localization
Mass spectrometry data are processed in analytical platforms (such as MaxQuant or Spectronaut), where experimental spectra are matched against theoretical peptide sequences in protein databases. Localization probabilities of phosphorylation sites are calculated, and only high-confidence sites are retained for downstream analyses.
5. Bioinformatic Interpretation
(1) Mapping kinase-substrate relationships (e.g., using KSEA and the PhosphoSitePlus database)
(2) Signaling pathway enrichment analysis (KEGG and Reactome)
(3) Differential phosphorylation site clustering, functional annotation, and network construction
(4) Integrated analysis with proteomic, transcriptomic, and metabolomic datasets
Phosphoproteomics vs. Proteomics vs. Transcriptomics: Unique Advantages
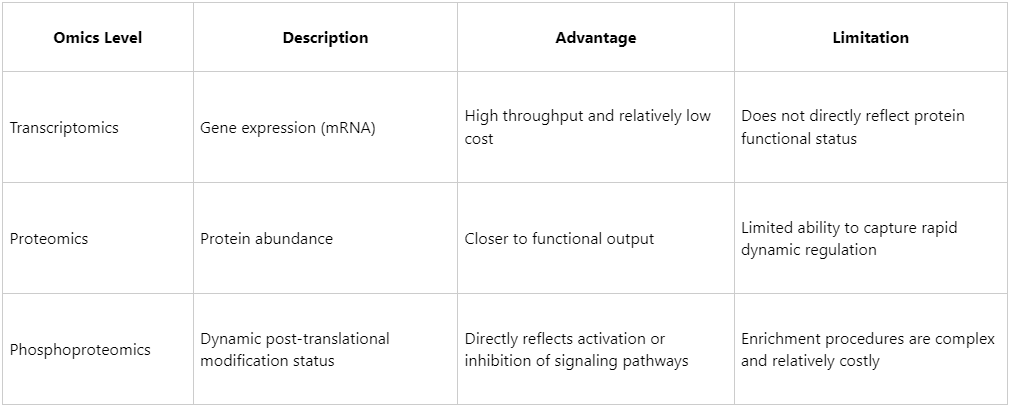
Phosphorylated protein analysis serves as a powerful tool for monitoring early signaling events and has indispensable value in research areas including oncology, inflammation, autophagy, and stem cell differentiation.
Phosphorylated protein analysis is not merely a technical approach but also a powerful framework for deciphering the dynamic regulatory logic of biological systems. Through this strategy, researchers can precisely identify critical nodes within signaling pathways, thereby guiding targeted therapeutic development, biomarker discovery, and mechanistic studies of disease. If you are encountering challenges in signaling pathway research or seeking to construct high-resolution cellular regulatory maps, MtoZ Biolabs stands ready to support your exploration of the phosphoproteome with advanced analytical solutions and technical expertise.
MtoZ Biolabs, an integrated chromatography and mass spectrometry (MS) services provider.
Related Services
How to order?







