How to Perform Proteomic Analysis on Paraffin-Embedded Tissue Samples?
-
Wash with xylene two to three times to dissolve paraffin.
-
Rehydrate sequentially using 100%, 95%, and 70% ethanol to remove residual organic solvents.
-
Finally, rinse with deionized water to eliminate remaining contaminants.
-
Extensive experience in handling FFPE specimens across multiple tissue types, including liver cancer, breast cancer, and colorectal cancer.
-
High-throughput mass spectrometry platforms supporting DDA, DIA, and PRM acquisition modes.
-
A specialized bioinformatics team capable of delivering customized analytical reports.
-
Capacity to undertake clinical collaborative projects and to support grant applications and scientific publications.
Formalin-fixed paraffin-embedded (FFPE) tissues are widely used in clinical pathology research, encompassing diverse disease types and extensive patient information, and therefore represent valuable resources for proteomics. However, protein crosslinking and chemical modifications introduced during formalin fixation and paraffin embedding pose significant technical challenges for proteomic analysis.
Key Challenges in FFPE Sample Proteomic Analysis
1. Protein Crosslinking and Denaturation
Formalin fixation induces formaldehyde-mediated crosslinks between proteins, particularly among lysine residues, resulting in denaturation, conformational alterations, and reduced solubility. These effects severely compromise subsequent protein extraction and enzymatic digestion efficiency.
2. Paraffin Embedding Interference
Paraffin is highly hydrophobic and therefore requires thorough removal; otherwise, residual paraffin can inhibit proteolytic digestion and contaminate the mass spectrometry system.
3. Sample Heterogeneity
FFPE sections are frequently derived from tumor tissues characterized by complex architectures and diverse cellular compositions. This necessitates stringent control of sample pretreatment to minimize potential bias.
Standard Workflow for FFPE Proteomic Analysis
MtoZ Biolabs has extensive experience supporting clinical projects involving tumors, inflammation, and autoimmune diseases. Based on large-scale FFPE sample analyses, the following optimized workflow is recommended:
1. Sectioning and Deparaffinization
(1) Sample preparation: Collect 5-10 µm FFPE sections and transfer them into RNase/DNase-free centrifuge tubes.
(2) Deparaffinization steps:
2. Antigen Retrieval and Protein Extraction
(1) Add a denaturing buffer, typically containing SDS, Tris-HCl, DTT, and related reagents, and incubate at high temperature (95°C) for an appropriate duration to reverse crosslinks.
(2) Protease inhibitors may be included to minimize proteolysis.
(3) Sonication or mechanical homogenization can be applied to enhance lysis efficiency and protein release.
3. Protein Quantification and Quality Control
(1) Determine protein concentration using the BCA or Bradford assay.
(2) Evaluate protein integrity and assess the presence of high-molecular-weight aggregates.
4. Digestion and Peptide Purification
(1) Apply SDS-PAGE or FASP procedures to remove detergents and other interfering substances.
(2) Digest proteins with trypsin under optimal conditions (37°C for 12–16 hours).
(3) Following digestion, purify peptides using C18 columns to eliminate salts and impurities.
5. LC-MS/MS Analysis
(1) Prior to analysis, peptide concentrations should be adjusted to an appropriate range to prevent column blockage.
(2) Acquire data using high-resolution mass spectrometers, such as Orbitrap Exploris or timsTOF Pro.
(3) Acquisition strategies, including DDA, DIA, or PRM, should be selected according to specific research objectives.
Data Analysis Strategy and Bioinformatics Interpretation
1. Data Search and Quantification
(1) Recommended software includes MaxQuant, Spectronaut, DIA-NN, and Proteome Discoverer.
(2) Databases: UniProt and RefSeq human protein databases.
(3) Quantification approaches may involve isobaric labeling strategies, such as TMT, or label-free methods.
2. Bioinformatics Analysis
(1) Identification of differentially expressed proteins (fold change combined with FDR correction).
(2) GO and KEGG enrichment analyses to investigate functional pathways.
(3) Construction of PPI networks and exploration of potential biomarkers.
Advantages and Application Prospects of FFPE Proteomics
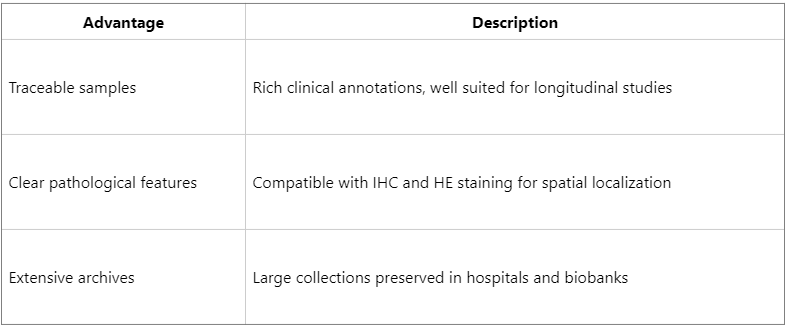
FFPE proteomics is progressively overcoming historical limitations and is becoming an essential technological foundation for precision medicine.
Advantageous Services of MtoZ Biolabs
Within the FFPE proteomics field, MtoZ Biolabs has established an integrated workflow spanning sample pretreatment, protein extraction, mass spectrometry analysis, and downstream data interpretation. Core strengths include:
Although FFPE specimens present technical difficulties, optimized preparation procedures combined with high-sensitivity mass spectrometry platforms make proteomic analysis fully achievable while providing substantial scientific and clinical value. For researchers seeking a reliable partner for FFPE proteomics services, please contact MtoZ Biolabs for professional support.
MtoZ Biolabs, an integrated chromatography and mass spectrometry (MS) services provider.
Related Services
How to order?







