How Does Formalin Fixation Influence Protein Digestion Efficiency?
-
The three-dimensional structure of proteins is altered.
-
Polypeptide chains become crosslinked, forming high-molecular-weight protein complexes.
-
Antigen epitopes and protease cleavage sites may be masked or structurally modified.
-
Cleavage sites may be chemically modified or sterically masked.
-
Crosslinked protein structures restrict enzymatic accessibility to substrate sites.
-
Trypsin catalytic activity is reduced or enzymatic reaction kinetics become constrained.
-
Excessively long fixation times (>24 h) significantly increase crosslinking density.
-
High formalin concentrations (>10%) can lead to more extensive chemical modifications.
-
Standard recommendation: 4% neutral buffered formalin with fixation times of 6-24 hours.
-
Dense tissues may slow fixative penetration, leading to heterogeneous fixation.
-
Both over-fixation and under-fixation can negatively influence subsequent proteolytic digestion.
-
Heat-induced antigen retrieval (e.g., 95-100°C) combined with high-pH buffers can help disrupt crosslinks.
-
Incomplete de-crosslinking prevents proteases from efficiently accessing target cleavage sites.
-
Organic solvents such as xylene are commonly used to completely remove paraffin.
-
Subsequent graded ethanol rehydration restores tissue hydrophilicity.
-
These steps ensure sufficient penetration of digestion buffers and proteases.
-
Buffers such as Tris-EDTA (pH 9.0) or citrate buffer (pH 6.0) are frequently used.
-
Water bath heating or pressure-assisted retrieval can partially disrupt crosslink bridges.
-
Proteases such as proteinase K, or enzyme combinations (e.g., trypsin + Lys-C), can improve digestion coverage.
-
MtoZ Biolabs has established optimized multi-protease digestion strategies suitable for difficult-to-digest samples such as FFPE tissues.
-
Digestion time can be extended to 12-24 hours, or sequential digestion rounds may be applied.
-
Care must be taken to avoid excessive autodigestion or non-specific degradation.
-
Advanced instruments such as Orbitrap Exploris and timsTOF Pro provide enhanced peptide detection capability.
-
MtoZ Biolabs employs high-resolution mass spectrometry platforms to enable accurate detection of low-abundance peptides.
-
Database searches can incorporate crosslink-related modifications and allow tolerance for non-specific cleavage.
-
Search engines supporting open-search strategies, such as MSFragger and Open-pFind, are particularly useful.
-
Customized FFPE-specific modification databases further improve the identification of atypical peptides.
Formalin-fixed paraffin-embedded (FFPE) tissue preservation has long been considered the gold standard for long-term storage of biological specimens and is widely used in pathology and clinical research. However, this preservation strategy also introduces substantial challenges for proteomic analysis, particularly with respect to proteolytic digestion efficiency.
Chemical Basis of Formalin Fixation and Its Impact on Protein Structure
1. Mechanism of Formaldehyde Crosslinking
The primary active component of formalin is formaldehyde. It reacts with amino acid residues in proteins (such as lysine, arginine, and cysteine), forming reversible or irreversible covalent crosslinks through methylene bridge formation. These crosslinking structures are highly stable.
As a result:
2. Effects on Protease (Especially Trypsin) Recognition Sites
Trypsin preferentially cleaves peptide bonds following lysine (K) and arginine (R) residues. However, when these sites undergo crosslinking:
Typical Effects of Formalin Fixation on Protein Digestion Efficiency
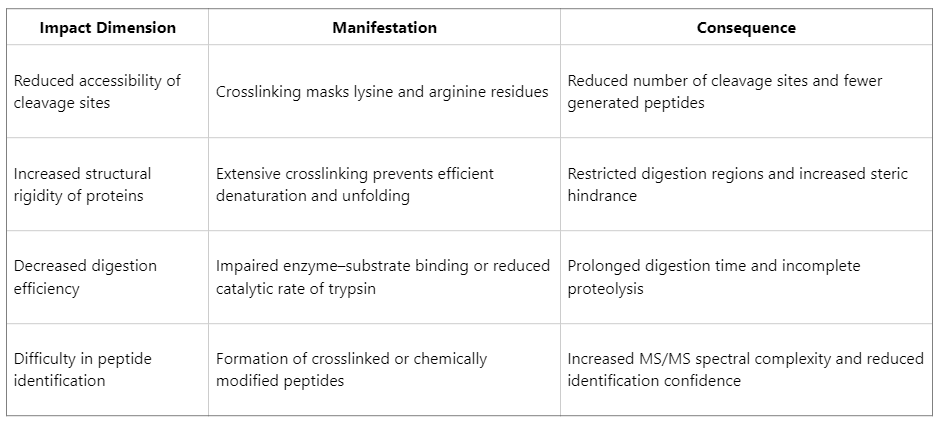
Key Factors Affecting Protein Digestion Efficiency
1. Fixation Time and Concentration
2. Tissue Type and Density
3. De-Crosslinking (Antigen Retrieval) Strategies
Strategies to Improve Protein Digestion Efficiency in FFPE Samples
1. Optimization of Deparaffinization and Rehydration Procedures
2. Application of Heat-Induced Epitope Retrieval (HIER)
3. Use of Auxiliary Proteases or Enzyme Mixtures
4. Extended Digestion Time or Multi-Step Digestion Strategies
Optimization Strategies at the Mass Spectrometry Level
1. High-Sensitivity Instrument Platforms
2. Optimization of Data Analysis Strategies
Although formalin fixation preserves tissue morphology and structural integrity, it significantly compromises proteolytic digestion efficiency and poses challenges for proteomic analysis. Understanding the underlying mechanisms and optimizing preprocessing workflows and digestion conditions are essential for overcoming bottlenecks in FFPE proteomics. In the field of FFPE-based mass spectrometry analysis, MtoZ Biolabs has established an integrated workflow covering deparaffinization, protein extraction, proteolytic digestion, and high-resolution mass spectrometry analysis. By addressing challenges such as inefficient proteolysis and modification interference, this workflow provides high-sensitivity and highly reproducible solutions, enabling researchers to maximize the proteomic information obtainable from valuable clinical samples.
MtoZ Biolabs, an integrated chromatography and mass spectrometry (MS) services provider.
Related Services
How to order?







