Histone Dimethylation Analysis
- Animal tissue ≥ 2 g
- Plant tissue ≥ 0.4 g
- Serum / Plasma ≥ 2 mL (plasma should be collected using EDTA as an anticoagulant)
- Urine ≥ 10 mL
- Cell ≥ 1 x 10⁸ cells
- Yeast / Microbial samples ≥ 0.4 g
- Protein samples: 2-5 mg (Standard tissue or cell lysis buffers can be used during protein extraction)
Histone dimethylation is a crucial post-translational modification that involves the addition of two methyl groups to specific lysine or arginine residues on histone proteins. This modification plays a pivotal role in regulating gene expression, chromatin structure, and overall cellular function. Histone dimethylation can either activate or repress gene expression, depending on the specific residue being modified and the context of other concurrent modifications.
Histone dimethylation is most commonly observed at key lysine residues, including H3K4, H3K9, H3K27, and H4K20. For example, dimethylation of H3K4 is often associated with transcriptional activation and is found in the promoters of actively expressed genes, while H3K9 dimethylation is linked to transcriptional repression and heterochromatin formation. This complex interplay between different methylation states underscores the importance of histone dimethylation in establishing and maintaining cellular identity, development, and response to environmental stimuli.
Recent research has highlighted the dynamic nature of histone dimethylation, revealing its involvement in various biological processes, including differentiation, DNA repair, and response to cellular stress. Additionally, dysregulation of histone dimethylation has been implicated in numerous diseases, particularly cancer, where alterations in the expression or activity of HMTs can lead to aberrant gene expression patterns.
Services at MtoZ Biolabs
MtoZ Biolabs utilizes Thermo Fisher's Q Exactive HF, Orbitrap Fusion, and Orbitrap Fusion Lumos mass spectrometry platforms, in combination with Nano-LC, to provide a comprehensive solution for histone dimethylation analysis. Simply provide your experimental objectives and send us your samples, and we will offer you a one-stop service from sample preparation and MS analysis to data analysis, delivering detailed identification and quantitative analysis of histone dimethylation modifications and their modification sites.
Compared to traditional proteomics analysis, the key to histone modification analysis lies in the isolation and purification of histone components, as well as mitigating the impact of N-terminal lysine on the analysis of histone post-translational modifications. Since histones are located in the cell nucleus, MtoZ Biolabs has developed a specialized platform for histone extraction and purification, drawing on existing literature and experimental experience. Cells are homogenized and subjected to low-speed centrifugation to obtain the nuclei. The nuclei are then further disrupted to release histones and DNA. High-salt buffer is used to remove impurities, and High-Performance Liquid Chromatography (HPLC) is employed to separate different histone components, achieving optimal purity for subsequent mass spectrometry analyses. Additionally, to eliminate the impact of N-terminal lysine, we utilize 2-3 different enzymes to enhance peptide coverage in mass spectrometry analysis during the digestion process. This multi-enzymatic digestion increases the coverage of histone peptides, preventing the loss of post-translational modification information due to unsuitable peptide lengths or low ionization efficiency.
Analysis Workflow
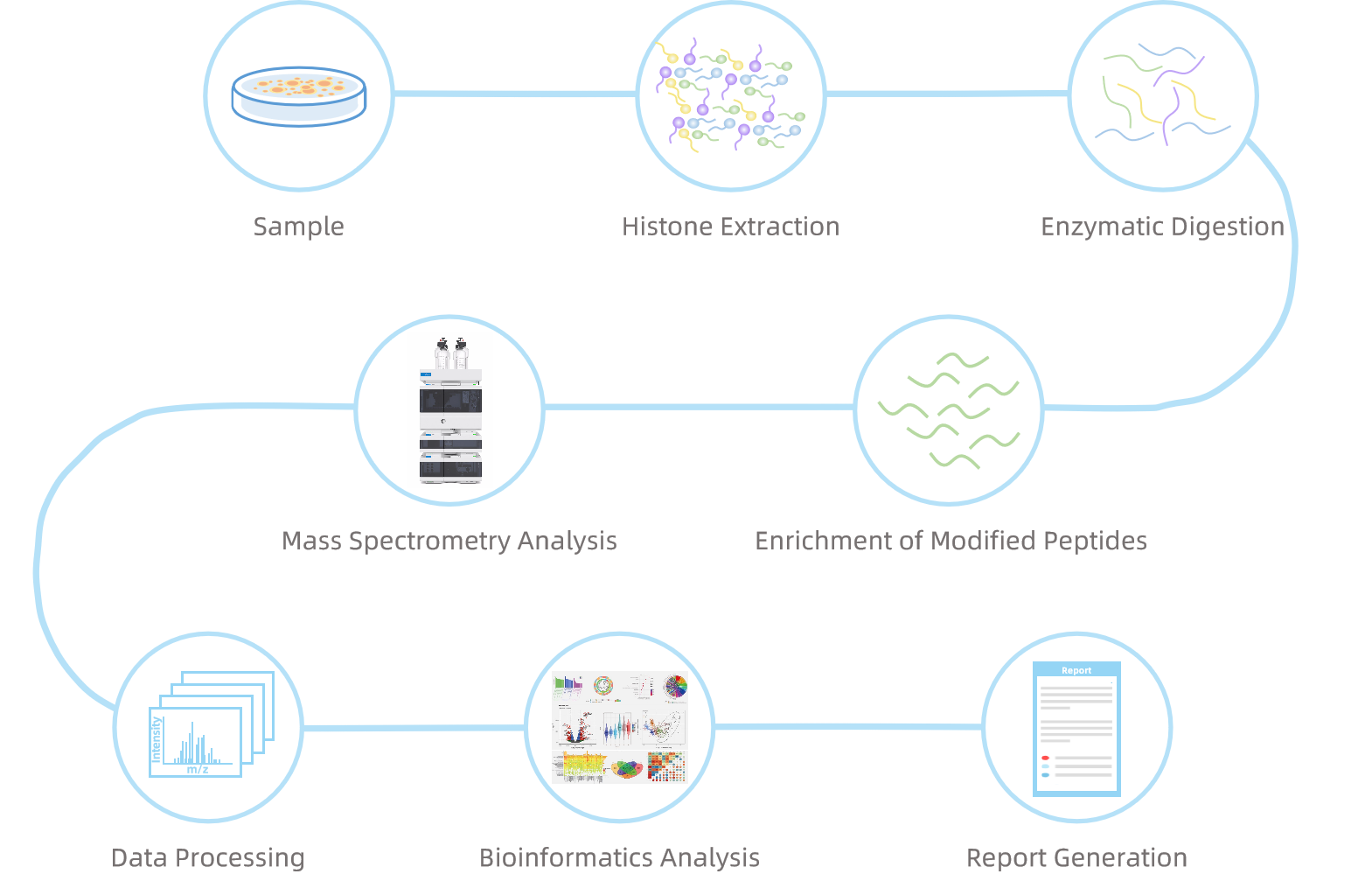
Service Advantages
1. Advance Analysis Platform
Mtoz Biolabs established an advanced histone dimethylation analysis platform, guaranteeing reliable, fast, and highly accurate analysis service.
2. One-Time-Charge
Our pricing is transparent, no hidden fees or additional costs.
3. High-Data-Quality
Deep data coverage with strict data quality control. AI-powered bioinformatics platform integrate all histone dimethylation analysis data providing clients with acomprehensive data report.
4. Optimized Extraction
An optimized histone extraction and purification protocols are employed to minimize contamination and ensure high purity.
Sample Submission Requirements
1. Sample Volume
2. Sample Preservation
Samples should be shipped with dry ice. Avoid repeated cycles of freezing and thawing
Note: We will conduct testing on the provided samples. The formal experiment will commence only after the samples pass the quality check.
Deliverables
1. Comprehensive Experimental Details (Materials, Instruments, and Methods, etc.)
2. Relevant Liquid Chromatography and Mass Spectrometry Parameters
3. Detailed Information on Histone Dimethylation Analysis
4. Raw Data
5. Mass Spectrometry Image
6. Custom Analysis Report
MtoZ Biolabs, an integrated Chromatography and Mass Spectrometry (MS) Services Provider, provides advanced proteomics, metabolomics, and biopharmaceutical analysis services to researchers in biochemistry, biotechnology, and biopharmaceutical fields. Our ultimate aim is to provide more rapid, high-throughput, and cost-effective analysis, with exceptional data quality and minimal sample consumption. If you are interested in our histone dimethylation analysis services, please feel free to contact us. We are dedicated to providing you with the best service.
MtoZ Biolabs, an integrated chromatography and mass spectrometry (MS) services provider.
Related Services
Histone Post-Translational Modification Analysis Service
How to order?







