HCP Antibody Development Service
Ideally, any technology for detecting Host Cell Protein (HCP) should: (i) detect protein concentrations across a wide dynamic range, from extremely low concentrations of individual impurity proteins to much higher concentrations of impurity proteins; (ii) track changes in HCP populations and concentrations throughout the entire bioprocess; (iii) simultaneously monitor and measure multiple protein analytes; and (iv) monitor and measure low concentration HCP even in the presence of target recombinant protein at high concentrations. Additionally, if a risk-based assessment of HCP is necessary, these methods should be capable of identifying HCP at any stage of the bioprocess. Employing anti-HCP antibodies for qualitative and quantitative detection and removal of HCPs is highly effective. Thus, the development of anti-HCP antibodies is of significant importance.
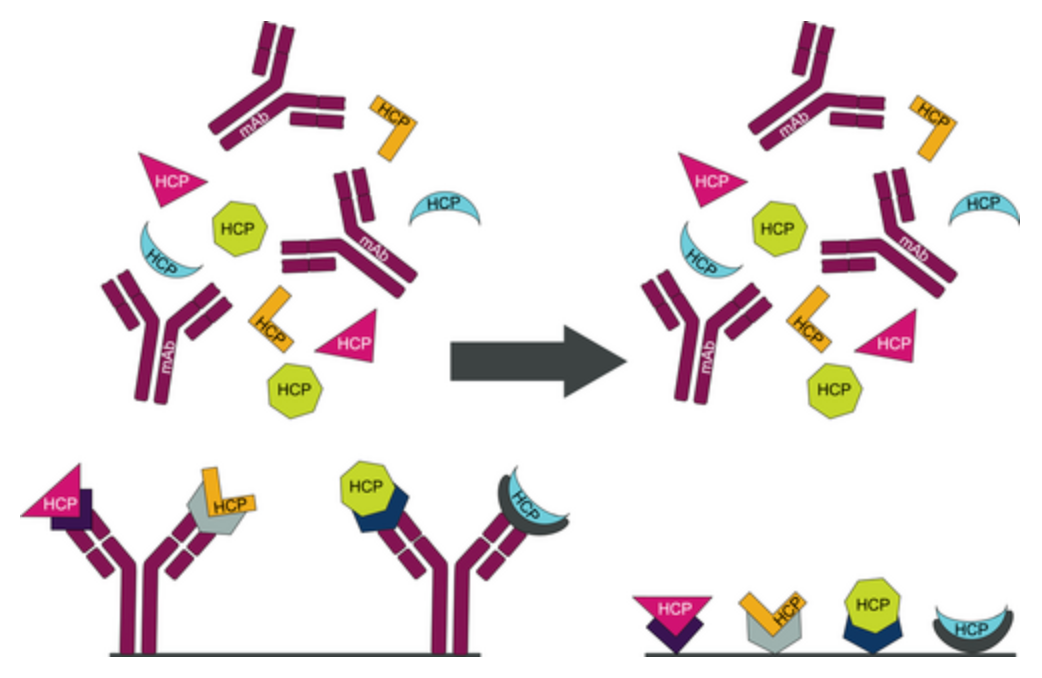
Figure 1. Multiple Methods for Capturing HCP (Antibody Capture (left) and Peptide Capture (right)) [1]
1. Identification of HCP Antigens
A variety of cell lines, including mammalian, insect, plant, and prokaryotic cells such as Escherichia coli, are employed for producing recombinant proteins for therapeutic purposes. Mammalian host cells are widely used in the biotechnology industry. These host cells express a multitude of proteins, potentially in the thousands, making the HCPs remaining in products highly heterogeneous, varying greatly in charge, hydrophobicity, and molecular weight. For instance, pIs range from ~3–11, with molecular weights from ~5 kDa to ~250 kDa for large proteins. During the purification process, the content of the product protein gradually increases, while the content of host impurity proteins significantly decreases, dropping from >10–90% to <1–100 ng/mg. Not only does the amount of host protein decrease, but the composition also changes.
Identification of HCP antigens involves two parts: comprehensive identification of all proteins expressed by the host cells and identification of HCPs predominantly remaining during the protein production process. By sequencing host cell genomes, a large portion of the extracellular proteome in cell culture fluids can be identified. Recently, around 3000 extracellular proteins have been identified, and eight different bioinformatics tools have been applied in the process of approximately 1000 proteins that may be secreted via classical and non-classical pathways. Techniques like allele labeling, spectral counting, and normalized spectral abundance factor (NSAF) are used for relative quantification of extracellular HCP in culture supernatants, while Mass Spectrometry Elevation (MSE, a tandem mass spectrometry technique with alternating high and low collision energies) is employed for absolute quantification.
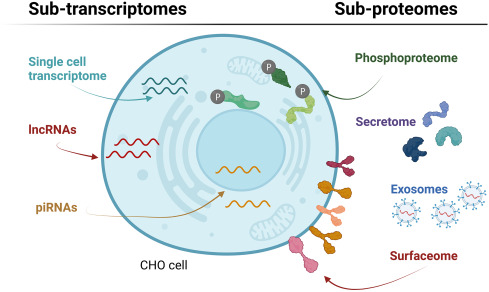
Figure 2. Subomics Characterization of CHO Cells [3]
2. Preparation of HCP Antigens
Total lysates or cell culture supernatants (such as from CHO cells) are utilized as immunogens to stimulate animals, typically goats or sheep, to produce antibodies. The resultant polyclonal antibodies encompass various IgGs against multiple CHO protein species, thus considered as multi-analyte anti-HCP antibodies. Protein western blot analysis with these antibodies demonstrates their ability to recognize diverse proteins. However, these antibodies' efficacy depends on the immunogenicity of HCPs within the animals that produce them. If the exogenous (host cell) proteins do not provoke or weakly provoke a reaction in the animal, the resultant polyclonal anti-HCP antibodies may be insufficient to detect those proteins. Additionally, if the host cell proteins elicit a reaction, but the resultant antibodies have low affinity or are produced in low quantities, detection of HCPs might be limited. Conversely, if HCPs are hyperimmunogenic, the antibodies produced may be abundant with high affinity, potentially leading to over-detection. Consequently, for HCP detection methods like ELISA and Western blotting that rely on these antibodies, detection and quantification may be distorted.
3. Animal Immunization
The immunogenicity strength of different antigens varies depending on their molecular weight, chemical active groups, stereostructure, physical shape, and diffusion speed. When immunizing animals, factors such as the dose of the antigen, the route of immunization, the number of immunizations, and the intervals between immunizations should be considered for their impact on the immunization outcome.
(1) The dose of the antigen: the immune dose of antigen depends on the type and route of antigen administration, the number of immunizations, and the species and immunization cycle of the recipient animal. Both excessive and insufficient doses can lead to immune tolerance. Moderate dose increases and extended intervals within the intermediate dose range can yield higher antibody titers. Typically, initial immunization doses range from 50-400μg for mice, 100μg-1mg for rats, and 200μg-1mg for rabbits, with booster doses being 1/5 to 2/5 of the initial dose. To prepare highly specific antisera, a low-dose antigen short-course immunization method can be used; if high-titer antisera are desired, it is appropriate to use a high-dose antigen long-course immunization method.
(2) Immunization routes, frequency and intervals: the immunization route, frequency, and intervals are selected based on antigenic properties, immunogenicity, and the animal's immune responsiveness. The route of antigen injection can be selected based on different antigens and experimental requirements, like intradermal, subcutaneous, intramuscular, intraperitoneal, intravenous, or lymph node injections. Typically, multiple injections are administered intradermally or subcutaneously at various sites such as the animal's back, footpad, around the lymph nodes, or behind the ear. The interval between two immunizations should be appropriately long, as too short an interval won't elicit an effective response, and too long an interval may lose the sensitization effect. Generally, the interval between the first and second immunizations is 10-20 days, with subsequent immunizations spaced 7-10 days apart. If adjuvants are used, the interval should be about 2 weeks. The total number of immunizations is usually 5-8 times.
(3) Application of adjuvants: adjuvants are used to enhance the immunogenicity of antigens, turning substances with no or weak immunogenicity into effective immunogens. For instance, when immunizing rabbits with soluble protein antigens, if adjuvants are used, the typical injection dose is about 0.5 mg/kg. Without adjuvants, the antigen dose should be increased by 10-20 times. The most commonly used adjuvants in animal experiments are Freund's Incomplete Adjuvant (FIA) and Freund's Complete Adjuvant (FCA). FIA is made by mixing the antigen with an oily substance (liquid paraffin or peanut oil) and then adding an emulsifier to create a water-in-oil emulsion. Freund's Complete Adjuvant is FIA with the addition of dead mycobacteria. For initial immunization, it is best to use Freund's Complete Adjuvant to stimulate a stronger immune response in the body. For subsequent immunizations, Freund's Incomplete Adjuvant is used. Soluble antigens often require the addition of adjuvants to enhance their immunogenicity and stimulate a stronger immune response in the body.
4. Antibody Purification
After establishing stable hybridoma cell lines, large-scale production of monoclonal antibodies (McAb) can commence. The main methods for McAb preparation include animal-induced ascites and in vitro culture.
(1) The animal-induced ascites method involves injecting an animal's peritoneal cavity with liquid paraffin or Freund's Incomplete Adjuvant. After one week, a suspension of hybridoma cells is injected into the animal's peritoneal cavity. Ascites can be collected in stages 10-14 days after inoculation. The collected ascites are centrifuged to remove cells, then inactivated (at 56°C for 30 minutes), followed by another round of centrifugation to obtain the supernatant. This method is simple, economical, and results in a high concentration of antibodies.
(2) The in vitro culture method involves cultivating hybridoma cells in a culture bottle. When the culture fluid changes color or cells begin to die excessively, the supernatant is collected, centrifuged to remove debris and cells, and then used.
Analysis Workflow
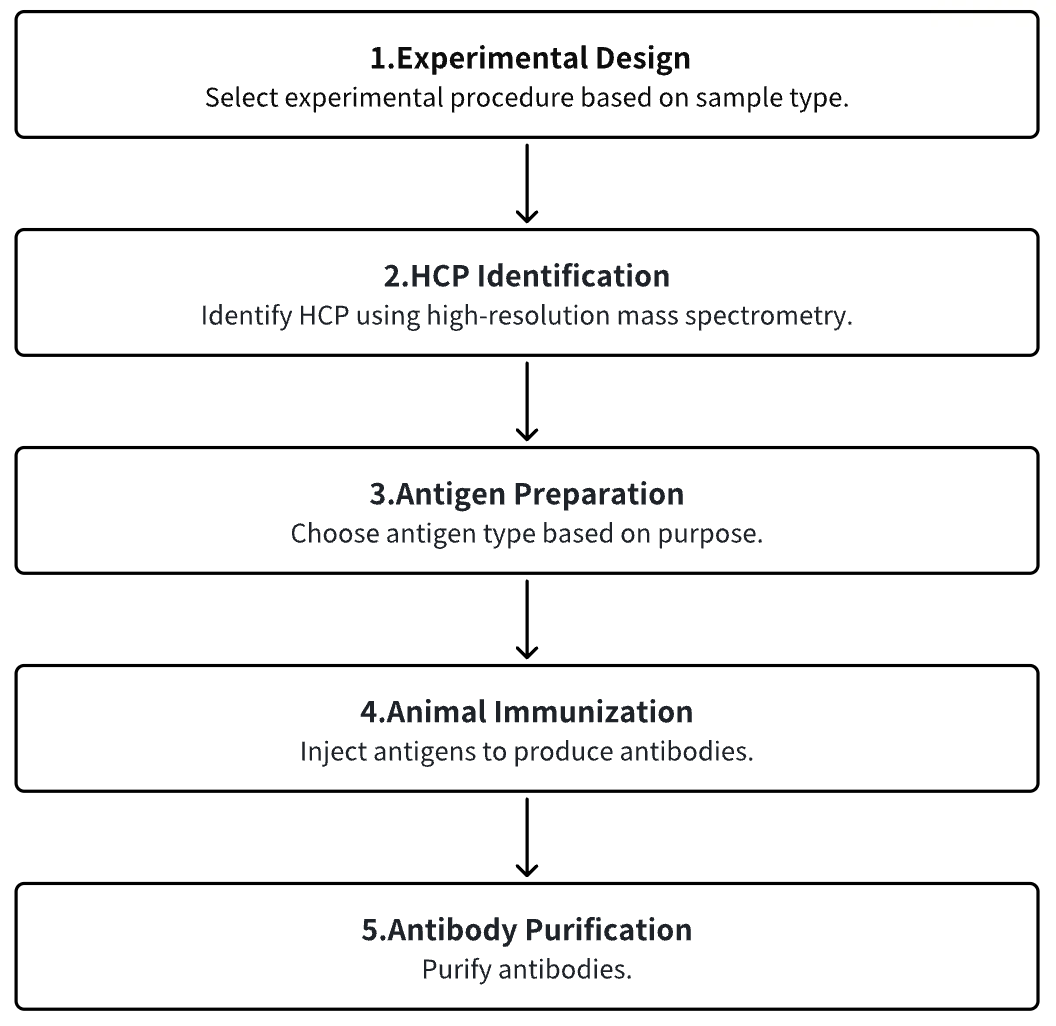
1. Experimental Procedure Determination
2. HCP Identification
3. Host Cell Antigen Preparation
4. Animal Immunization
5. Antibody Purification
Service Advantages
1. High Reliability Identification and Characterization of HCP
2. Multiple Antigen Preparation Platforms
3. Comprehensive and Strict Immunization Procedures
4. Effective Antibody Detection Systems and Extensive Detection Experience
Sample Results
1. Identification and Characterization of CHO HCPs Co-Purified with Monoclonal Antibodies
HCPs are impurities in biopharmaceutical production that can affect the safety and effectiveness of therapeutic monoclonal antibodies (mAbs). Thus, understanding the interactions between HCP impurities and mAbs is key to efficient downstream process design. There are studys have employed two methods to identify co-purified HCP subgroups: incubation of purified mAbs with media from mock-transfected CHO cells, and immobilization of mAbs on chromatographic media followed by incubation. Total HCP levels were measured using a CHO HCP ELISA, and changes in CHO HCPs were detected using 2-DE and LC-MS/MS. Results revealed higher HCP contents in protein A product pools compared to controls, with varying HCP contents across different protein A purification fractions. LC-MS/MS identified most HCPs in three protein A product pools, showing overlap with HCPs in eluates from columns with different mAbs immobilized. Additionally, the HCPs associated with mAbs predominantly played catalytic roles, suggesting mAbs can bind to both common and unique HCPs.
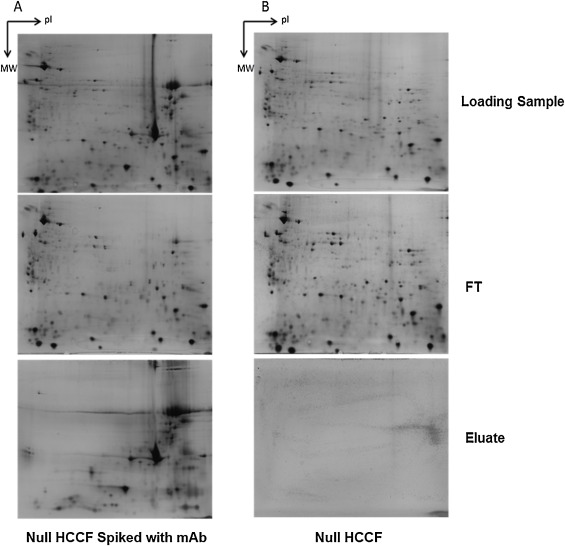
Figure 3. Representative 2-DE Silver Staining Images of Components During Protein A Affinity Chromatography Purification Process [4]
2. Characterization of HCPs Co-Eluted with mAbs in Protein A Purification
Protein A chromatography is commonly the initial step in purifying mAb biopharmaceuticals expressed in mammalian tissue culture cells. The goal of this step, and subsequent chromatographic processes, is partly to remove HCPs and other impurities. Understanding the retention mechanism of HCP subsets in this phase is crucial for mAb process developers, as it can inform targeted HCP removal strategies. However, specific information on HCPs co-purified with mAbs is limited. Research has employed 2D-LC-HDMS to comprehensively compared HCP subgroups associated with 15 different mAbs during protein A chromatography. It was found that most CHO HCPs binding and eluting with mAbs were common across the studied antibodies. In Protein A purifications, only a small portion of the total mAb HCP content (on average about 10%) elutes with the specific antibodies. The abundance of these HCPs in the culture fluid and their interaction capability with mAbs were key factors determining their prevalence in Protein A eluates. The study also investigated potential binding fragments between HCPs and mAbs by co-purifying them with individual Fc and (Fab′)2 antibody fragments.
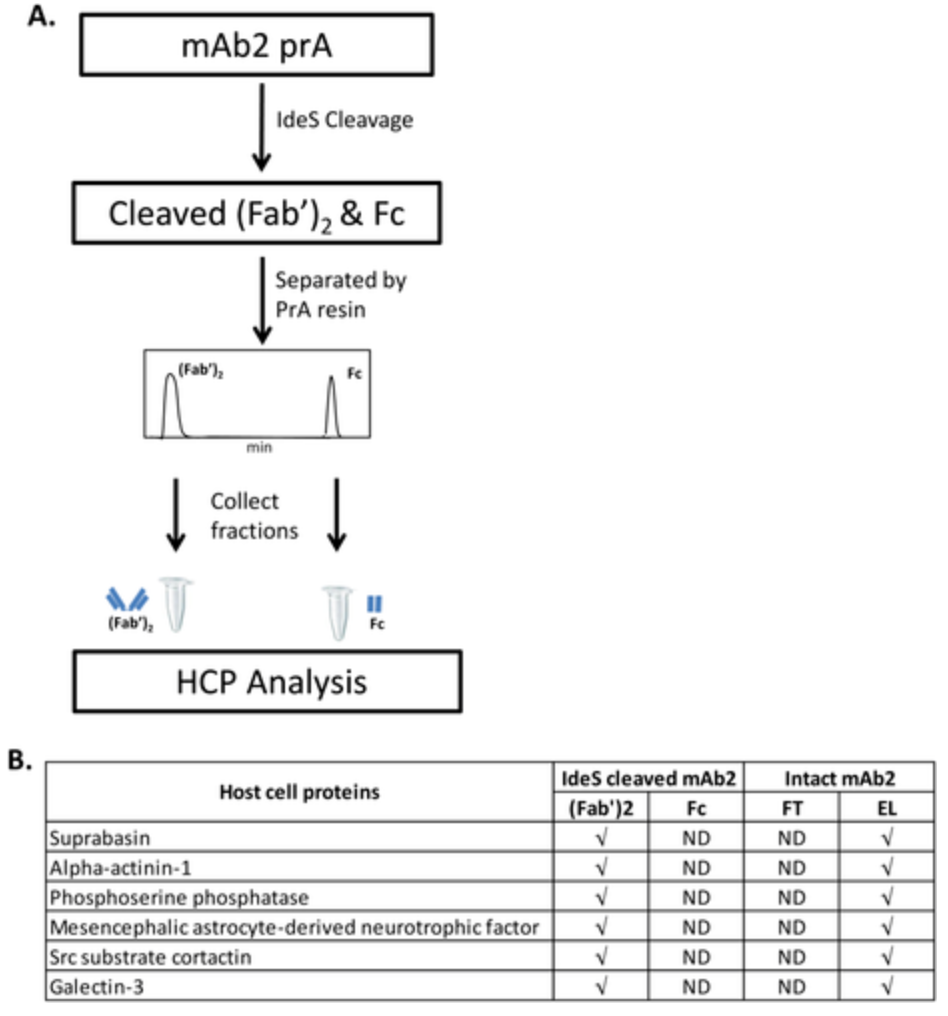
Figure 4. Detection of MAb-Specific HCP in the Fab Domain of mAb2 [5]
3. Quantitative Analysis of HCPs in CHO and Vero Cell Line Biopharmaceuticals Using Capillary Western Blot (Wes)
Quantifying HCPs is critical in manufacturing many biologics and vaccines. Common methods include enzyme-linked immunosorbent assay (ELISA), mass spectrometry (MS), and other orthogonal assays. Prior to using these techniques, it is necessary to evaluate key reagents, such as assessing the HCP coverage of antibody. HCP coverage percentage is typically determined by denaturing 2D Western blotting. However, ELISA only measures HCP quantities in their native state. Studies linking reagents validated by 2D Western blotting to sufficient coverage in final ELISAs are limited. The newly developed Wes technique by ProteinSimple allows semi-automated, streamlined protein separation, blotting, and detection. Wes is similar to traditional Western blotting (WB) but has the added advantage of quantification. One research outlined the Wes approach that connects 2D Western blotting coverage with final ELISAs for more effective HCP quantification. This study describes the development of Wes method for quantitatively evaluating HCPs in Vero and CHO cell lines. As expected, CHO HCP quantities decreased with increasing sample purity. The method identified similar quantities of Vero HCPs in both denatured (WES) and native (ELISA) detection formats. Additionally, this method can quantify the coverage of anti-HCP antibody reagents used in commercial HCP ELISA kits.
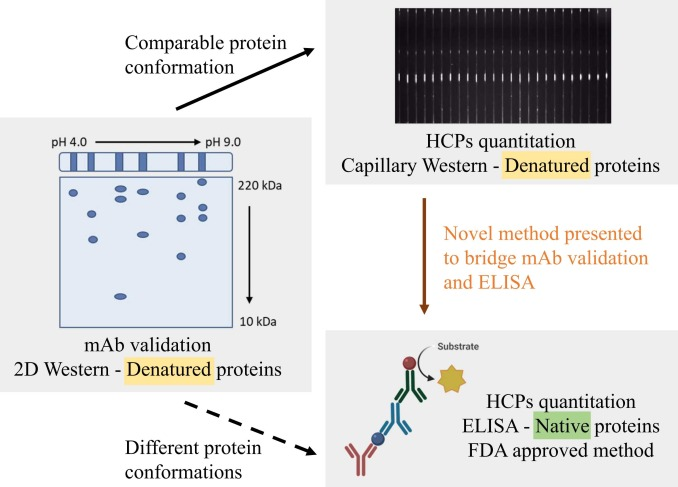
Figure 5. Modified Wes for mAbValidation [6]
Sample Submission Requirements
1.Minimize Impurity Contamination
Services at MtoZ Biolabs
1. Complete Experimental Procedure
2. Relevant Instrument Parameters
3. Original Experimental Data
4. Data Analysis Submission
Applications
1. Biopharmaceutical Production Process Optimization
Since the composition and concentration of HCP in specific drug products may be influenced by upstream and downstream processing conditions, variations in the process can affect the HCP profile. Consequently, monitoring HCP can be utilized to optimize the production process.
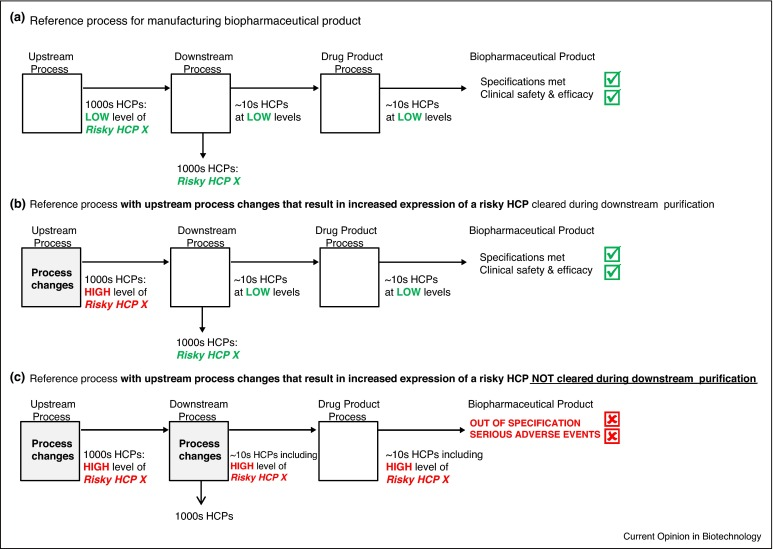
Figure 6. Modifications to the manufacturing process (a) can alter the HCP profile through changes in (b) upstream processes and (c) downstream processes, thereby negatively impacting product quality. [7]
FAQ
Q1: What are the types of HCP? What are their negative impact on mAb production and quality?
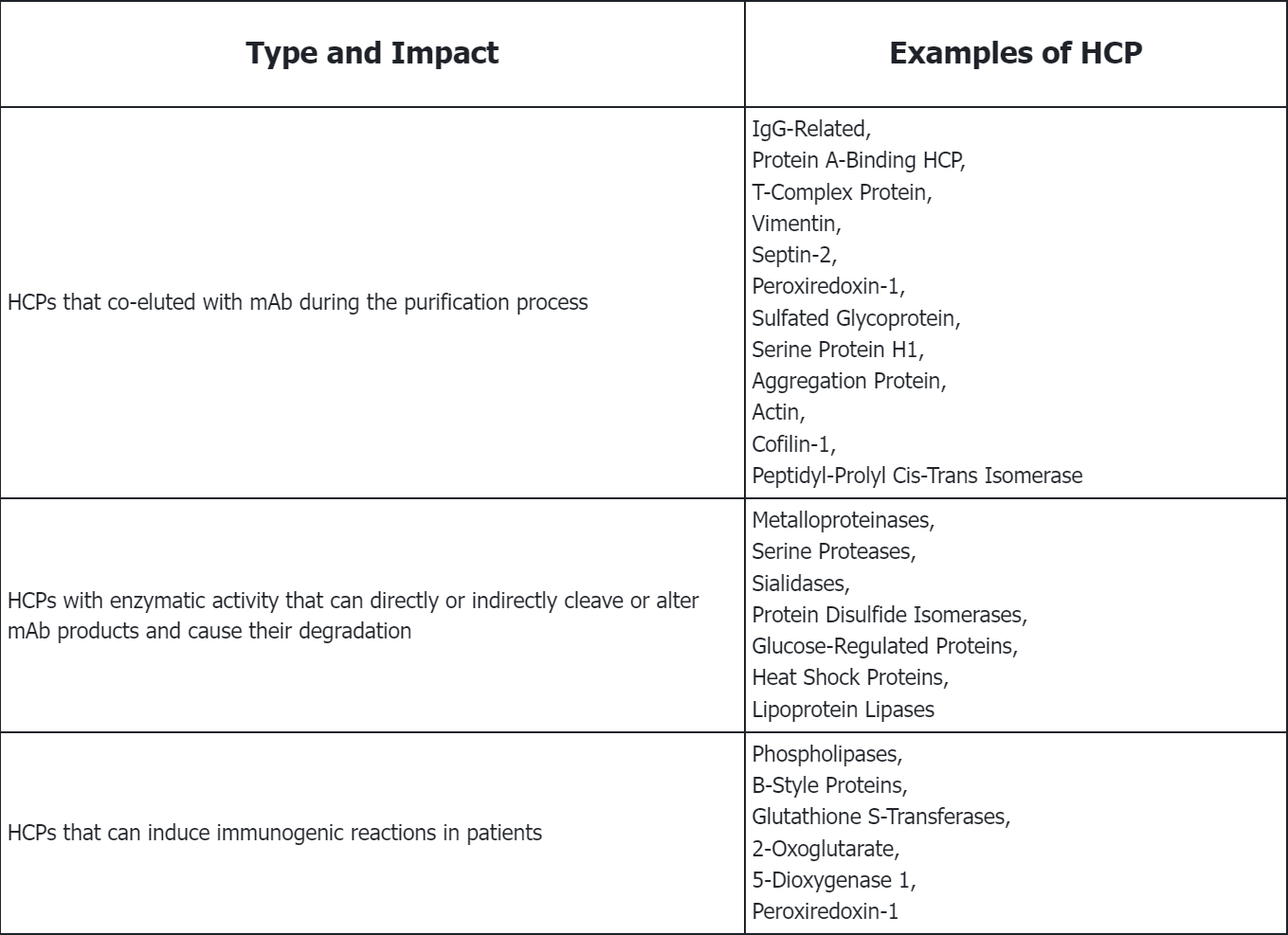
References
[1] Tuameh A, Harding SE, Darton NJ. Methods for addressing host cell protein impurities in biopharmaceutical product development. Biotechnol J. 2023 Mar;18(3):e2200115. doi: 10.1002/biot.202200115. Epub 2022 Dec 13. PMID: 36427352.
[2] Haltaufderhyde K, Roberts BJ, Khan S, Terry F, Boyle CM, McAllister M, Martin W, Rosenberg A, De Groot AS. Immunoinformatic Risk Assessment of Host Cell Proteins During Process Development for Biologic Therapeutics. AAPS J. 2023 Sep 11;25(5):87. doi: 10.1208/s12248-023-00852-z. Erratum in: AAPS J. 2023 Dec 19;26(1):6. PMID: 37697150.
[3] Jerabek T, Keysberg C, Otte K. The potential of emerging sub-omics technologies for CHO cell engineering. Biotechnol Adv. 2022 Oct;59:107978. doi: 10.1016/j.biotechadv.2022.107978. Epub 2022 May 13. PMID: 35569699.
[4] Liu X, Chen Y, Zhao Y, Liu-Compton V, Chen W, Payne G, Lazar AC. Identification and characterization of co-purifying CHO host cell proteins in monoclonal antibody purification process. J Pharm Biomed Anal. 2019 Sep 10;174:500-508. doi: 10.1016/j.jpba.2019.06.021. Epub 2019 Jun 18. PMID: 31234041.
[5] Zhang Q, Goetze AM, Cui H, Wylie J, Tillotson B, Hewig A, Hall MP, Flynn GC. Characterization of the co-elution of host cell proteins with monoclonal antibodies during protein A purification. Biotechnol Prog. 2016 May;32(3):708-17. doi: 10.1002/Epub 2016 May 6. PMID: 27073178.
[6] Valente KN, Levy NE, Lee KH, Lenhoff AM. Applications of proteomic methods for CHO host cell protein characterization in biopharmaceutical manufacturing. Curr Opin Biotechnol. 2018 Oct;53:144-150. doi: 10.1016/j.copbio.2018.01.004. Epub 2018 Feb 3. PMID: 29414072.
MtoZ Biolabs, an integrated chromatography and mass spectrometry (MS) services provider.
Related Services
How to order?







