Cysteine Redox Proteomics Service
Cysteine Redox Proteomics Service is a mass spectrometry-based post-translational modification proteomics service designed to detect and quantify redox modifications of cysteine residues in proteins. Cysteine residues, due to the chemical reactivity of their thiol group (-SH), are primary targets for redox modifications. In biological systems, cysteine thiol groups can undergo various oxidation modifications, such as S-thiolation, S-glutathionylation, S-sulfenylation, and disulfide bond formation. These modifications not only play crucial roles in regulating protein function, stability, and interactions but are also closely associated with cellular signaling, oxidative stress responses, and numerous diseases including cancer, neurodegenerative disorders, and cardiovascular diseases. Utilizing high-sensitivity mass spectrometry combined with strategies such as thiol enrichment and selective chemical labeling, MtoZ Biolabs offers Cysteine Redox Proteomics Service systematically identifying the redox state, modification types, sites, and dynamic changes of cysteine thiols in proteins, providing high-quality experimental data supporting research on oxidative stress mechanisms, disease molecular mechanisms, and drug target discovery.
Analysis Workflow
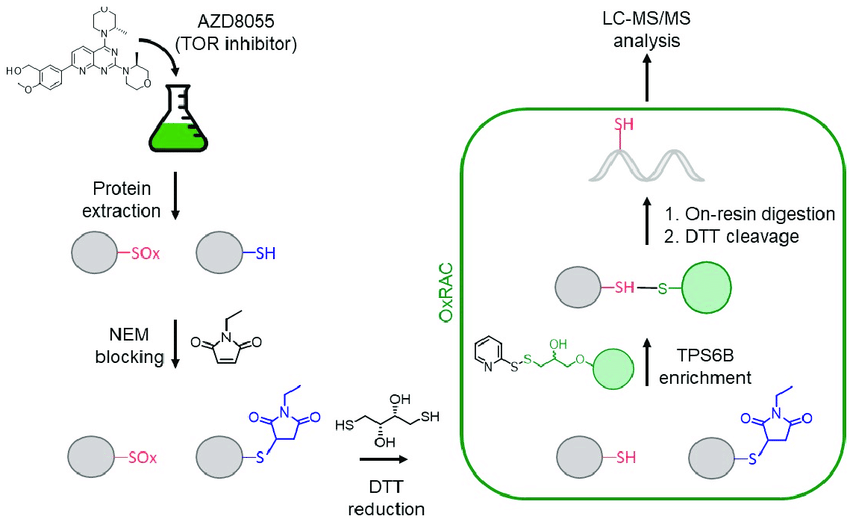
Ford M M. et al. Cells. 2019.
Figure 1. Example Workflow of Cysteine Redox Proteomics.
1. Sample Preparation and Treatment
Extract proteins and treat with a reducing agent, followed by labeling with thiol-specific chemical reagents to distinguish different cysteine redox states.
2. Enrichment and Isolation of Thiol-Modified Peptides
Use specific biotinylation strategies or thiol-affinity capture resins to enrich targeted modified peptides. Remove non-specifically bound products to obtain high-purity thiol-modified peptide samples.
3. LC-MS/MS Analysis
Separate peptides using nano-liquid chromatography (nano-LC) and detect peptide ions using high-resolution mass spectrometry, acquiring parent-ion and fragment-ion spectra.
4. Data Processing and Quantification
Identify and quantify modified peptides using specialized proteomics analysis software. Conduct functional annotation, pathway analysis, and network construction to comprehensively interpret the biological significance of identified modifications.
Applications
Applications of Cysteine Redox Proteomics Service include, but are not limited to:
Redox Mechanism Studies in Disease
Analyzing redox regulatory networks in oxidative stress-related diseases to uncover key pathological mechanisms.
Investigating the relationship between cellular metabolic reprogramming and redox homeostasis to identify potential therapeutic targets.
Oxidative Stress and Signal Transduction Analysis
Identifying dynamically changing redox-sensitive proteins under oxidative stress conditions and elucidating their roles in signaling pathways.
Deepening understanding of redox regulation in the cell cycle, gene expression, and inflammatory responses.
Drug Development and Target Validation
Quantitatively assessing changes in redox modifications under potential drug treatments to evaluate drug effects on target proteins.
Supporting the screening and validation of antioxidant drugs or modulators and their mechanisms of action.
Molecular Biomarker Discovery
Identify redox modification sites associated with disease progression and provide candidate biomarkers for early diagnosis or prognostic assessment.
Integrating modification-site information into multi-omics analyses to develop novel diagnostic tools.
Services at MtoZ Biolabs
MtoZ Biolabs, an integrated Chromatography and Mass Spectrometry (MS) Services Provider, provides advanced proteomics, metabolomics, and biopharmaceutical analysis services to researchers in biochemistry, biotechnology, and biopharmaceutical fields. Our ultimate aim is to provide more rapid, high-throughput, and cost-effective analysis, with exceptional data quality and minimal sample consumption. Leveraging high-resolution mass spectrometry, MtoZ Biolabs provides Cysteine Redox Proteomics Service dedicated to identifying cysteine-modified proteins and their modification sites, as well as quantitatively analyzing their dynamic changes. By dissecting protein redox states, we support in-depth studies on oxidative stress response mechanisms, elucidate disease-related molecular mechanisms, and offer scientific evidence for discovering novel drug targets.
FAQ
Q. How can enrichment efficiency and specificity for thiol modifications be improved?
1. Optimize reagent selection: Use highly specific thiol-labeling reagents (e.g., N-ethylmaleimide or iodoTMT) to selectively label cysteine residues, avoiding nonspecific reactions caused by excess reagent.
2. Improve enrichment strategies: Employ capture materials with greater affinity, such as biotinylated thiol-capture reagents, combined with multiple enrichment rounds to enhance the recovery of target modified peptides.
3. Strictly control experimental conditions: Avoid nonspecific oxidation during experiments by maintaining appropriate pH levels and salt concentrations to reduce background interference.
4. Incorporate separation techniques: Introduce a preliminary high-performance liquid chromatography (HPLC) step before enrichment to remove highly abundant interfering components, thus increasing the specificity of subsequent enrichment.
Q. How should experimental conditions be designed to capture dynamic changes in thiol modifications?
1. Select appropriate time points: Design experiments to include multiple sampling points following cellular stimulation, drug treatment, or oxidative stress exposure to capture the full progression of modification dynamics from initial states onward.
2. Control stimulus dosage: Apply gradient concentrations of stimulating agents or oxidants (e.g., hydrogen peroxide) to cells or tissues, thereby analyzing concentration-dependent changes and clarifying the relationship between dosage and modification levels.
3. Introduce internal reference controls: Add stable internal reference proteins or standards to ensure that observed differences in modifications across conditions genuinely reflect biological effects rather than technical variations.
4. Optimize enrichment and mass spectrometry protocols: Keep consistent enrichment procedures and mass spectrometry parameters across all time points to minimize technical variability, ensuring that dynamic changes accurately represent the biological state of modifications.
Deliverables
1. Comprehensive Experimental Details
2. Materials, Instruments, and Methods
3. Total Ion Chromatogram & Quality Control Assessment (project-dependent)
4. Data Analysis, Preprocessing, and Estimation (project-dependent)
5. Bioinformatics Analysis
6. Raw Data Files
Case Study
This study utilized Cysteine Redox Proteomics to investigate the effects of the thioredoxin reductase 1 (TrxR1) inhibitor Auranofin on MYC-overexpressing high-grade serous ovarian cancer (HGSOC) cells. The results demonstrated that Auranofin treatment induced oxidation of cysteine residues in multiple proteins, particularly key enzymes in the glycolytic pathway, such as glyceraldehyde-3-phosphate dehydrogenase (GAPDH). Oxidation of these residues significantly inhibited enzyme activity, thereby suppressing glycolysis. Under conditions of glycolytic inhibition, MYC-overexpressing HGSOC cells shifted their metabolic dependence toward glutamine metabolism to maintain survival. Further experiments revealed that combined treatment with Auranofin and the glutaminase (GLS1) inhibitor CB-839 markedly enhanced antitumor efficacy. This study highlights the potential therapeutic strategy of simultaneous TrxR1 and GLS1 inhibition for treating MYC-driven HGSOC patients.
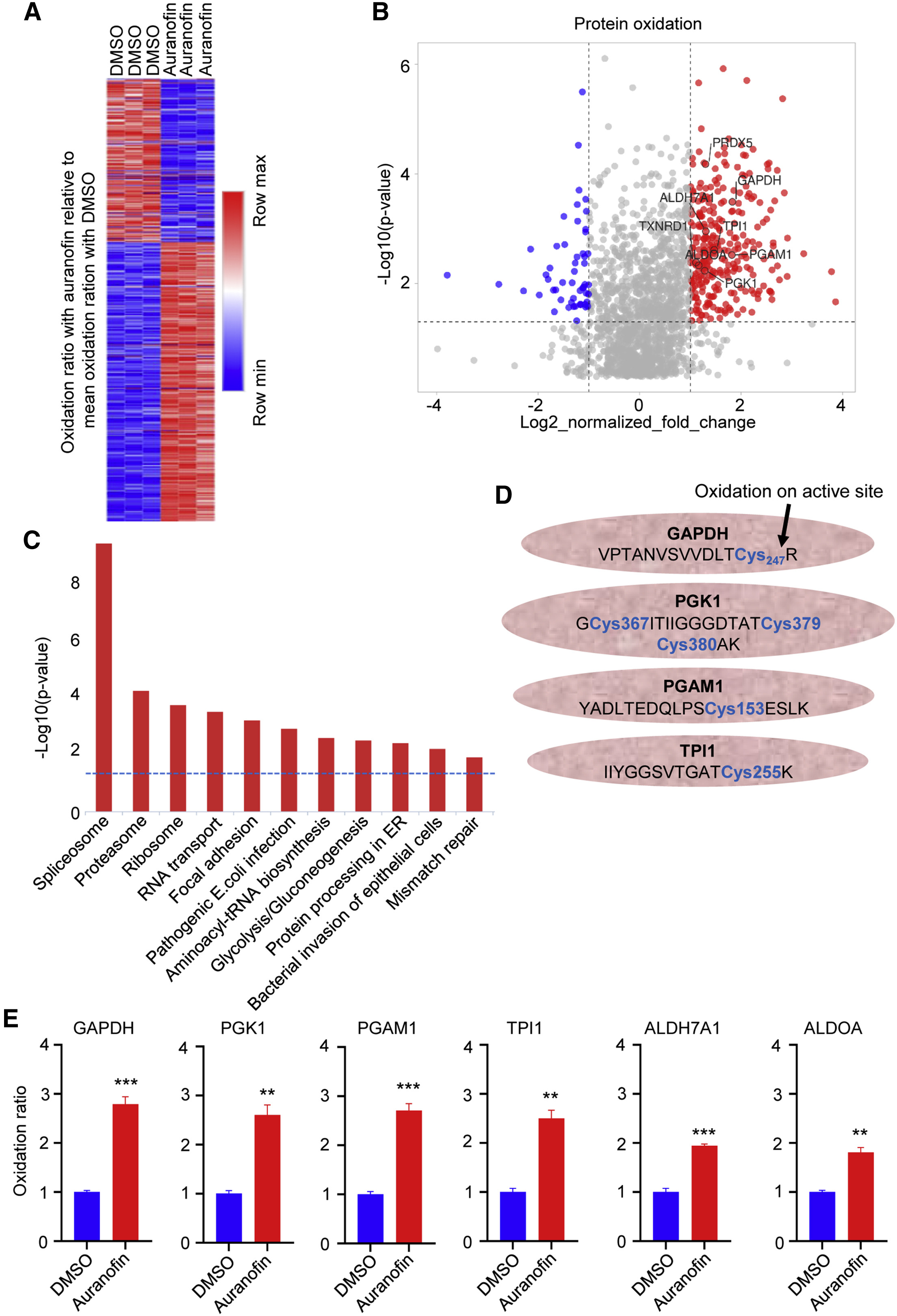
Raninga P V. et al. Molecular Therapy. 2023.
Figure 2. The redox proteome of MYC-high HGSOC cells after auranofin treatment.
MtoZ Biolabs, an integrated chromatography and mass spectrometry (MS) services provider.
Related Services
Post-Translational Modifications Proteomics Service
How to order?







