ChIRP-Seq Service
-
Cultured cells: ≥ 1 × 107 cells
-
Fresh or Frozen Tissues: ≥ 50 mg
MtoZ Biolabs provides a comprehensive ChIRP-Seq Service for systematic analysis of long noncoding RNA mediated interactions with chromatin and associated protein complexes. This service enables integrated investigation of lncRNA DNA binding sites, lncRNA associated proteins, and lncRNA driven regulatory mechanisms at genome-wide scale.
What Is ChIRP-Seq?
Long noncoding RNAs (lncRNA) play essential roles in transcriptional regulation chromatin remodeling nuclear organization and epigenetic control. Unlike messenger RNAs many lncRNAs function as molecular scaffolds or guides that coordinate interactions between chromatin and regulatory proteins. Understanding where a lncRNA binds in the genome and which proteins associate with it is critical for deciphering its biological function.
Chromatin Isolation by RNA Purification followed by sequencing commonly known as ChIRP-Seq is a targeted strategy developed to capture endogenous lncRNAs together with their associated DNA and protein partners. By directly isolating lncRNA centered chromatin complexes from cells ChIRP-Seq enables simultaneous analysis of RNA DNA and protein interactions in a native cellular context. ChIRP-Seq has become a foundational approach for functional lncRNA studies especially in transcriptional regulation epigenetics cancer biology and developmental biology where RNA mediated chromatin interactions play central roles.
Principle of ChIRP-Seq
ChIRP-Seq relies on sequence-specific hybridization of biotinylated antisense DNA probes tiled across the target RNA transcript. Short complementary probes are designed at regular intervals along the RNA and divided into two independent pools based on alternating positions to ensure specificity and reproducibility.
After hybridization, RNA-centered complexes are captured using streptavidin-coated magnetic beads, allowing co-enrichment of DNA and proteins associated with the target RNA. The enriched DNA and protein fractions are subsequently separated and analyzed by next-generation sequencing and mass spectrometry, respectively, enabling integrated mapping of RNA-DNA and RNA-protein interactions.
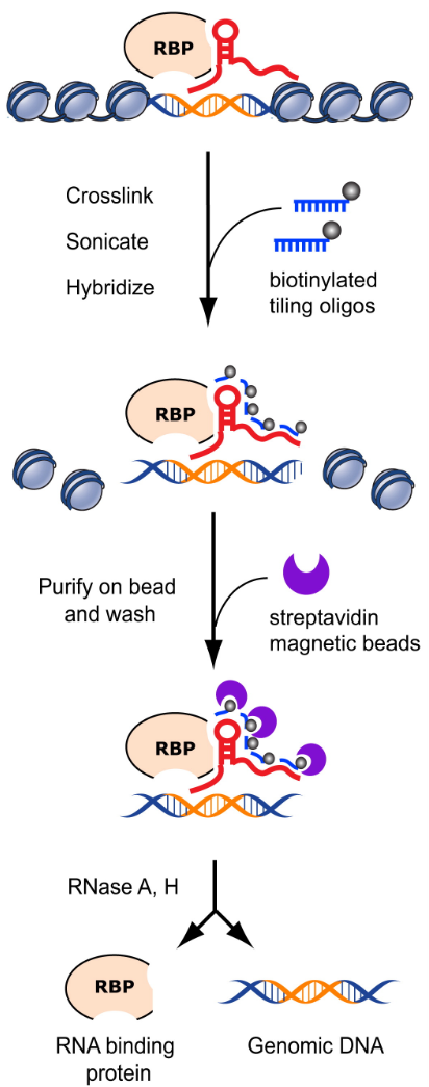
Chu, C. et al. Mol Cell. 2011.
Figure 1. Principle of ChIRP
Service at MtoZ Biolabs
MtoZ Biolabs provides integrated ChIRP-based solutions for comprehensive analysis of nucleic acids and proteins associated with target lncRNAs .
💠ChIRP-Seq by NGS
Genome-wide identification of DNA regions associated with the target lncRNA or circRNA.
💠ChIRP-qPCR
Targeted validation of specific DNA or RNA interactions with the target lncRNA or circRNA.
💠ChIRP-MS
Proteome-wide identification of proteins interacting with the target lncRNA or circRNA.
💠ChIRP-Western Blot
Targeted confirmation of candidate protein interactions with the target lncRNA or circRNA.
Workflow of ChIRP-Seq Service
1. Experimental design and probe set optimization
2. Crosslinking and chromatin preparation
3. Hybridization of biotinylated probes to target lncRNA
4. Affinity capture of lncRNA centered complexes
5. Stringent washing to reduce nonspecific background
6. Separation of DNA and protein fractions
7. DNA analysis by qPCR or next-generation sequencing (NGS)
8. Protein analysis by mass spectrometry, SDS-PAGE or Western blotting
9. Data processing and integrated biological interpretation
Why Choose MtoZ Biolabs
☑️High Data Quality
Optimized experimental workflows and rigorous quality control ensure reliable and reproducible results.
☑️Advanced Analytical Platforms
Integrated sequencing and mass spectrometry platforms support high-sensitivity detection of RNA, DNA, and protein interactions.
☑️Extensive Project Experience
Proven expertise across diverse RNA species and experimental designs supports confident execution of complex ChIRP studies.
☑️Customized Study Design
Experimental strategies are tailored to RNA characteristics, sample types, and specific research objectives.
☑️Efficient Project Turnaround
Streamlined operations and coordinated workflows enable timely delivery without compromising data quality.
Applications of ChIRP-Seq Service
1. Disease Research
Map RNA-centered chromatin and protein interaction networks to clarify how lncRNAs or other ncRNAs regulate pathogenic gene programs in cancer, neurological disorders, immune diseases, infectious disease, and metabolic disorders.
2. Drug Discovery and Development
Support target identification and validation by linking candidate RNAs to specific genomic loci and protein partners, enabling mechanism-of-action studies, biomarker development, and evaluation of RNA-targeting modalities.
3. Basic Research in Gene Regulation and Epigenetics
Define RNA–DNA binding landscapes and RNA-associated protein complexes to investigate transcriptional control, chromatin remodeling, nuclear organization, and RNA-driven regulatory pathways.
4. Functional Annotation of lncRNAs
Assign biological roles to uncharacterized RNAs by discovering their chromatin targets and interaction partners, guiding downstream perturbation and validation studies.
5. Comparative and Condition-Dependent Interaction Studies
Compare RNA interaction profiles across cell types, developmental stages, treatments, or stress conditions to uncover dynamic RNA-mediated regulatory mechanisms.
FAQ
Q1: What types of samples are suitable?
For reliable ChIRP-Seq enrichment and sequencing, we typically recommend the following sample inputs.
For unusual sample types or low-yield materials, please contact MtoZ Biolabs in advance for customized preparation guidance.
Q2: How should I prepare my samples?
1. Collect and process samples consistently across groups and minimize RNA degradation risk
2. Keep handling cold, use RNase-free consumables, and avoid introducing RNase contamination
3. Snap-freeze samples promptly, store at -80°C, and avoid repeated freeze-thaw cycles
4. Ship on dry ice with secure packaging and clear labels for sample ID, group, and replicate
5. Provide your research objective, target RNA information (sequence or transcript ID), species, and group design to align probe design, controls, and analysis
For more information, please refer to Sample Submission Guidelines for Proteomics and Sample Submission Guidelines for Metabolomics.
Q3: What is the service general workflow?
Q4: What data formats are provided?
1. Raw data: FASTQ (NGS), vendor MS raw files and optional mzML (MS)
2. Processed tables: XLSX or CSV (peak or target lists, enrichment summaries, qPCR results, protein IDs, and quantification when included)
3. Alignments and tracks: BAM with index files and BigWig tracks (for ChIRP-Seq projects)
4. Images and figures: TIFF or PNG plus PDF (QC plots, summary figures, and western blot images)
5. Reports and methods: PDF (assay design, controls, key parameters, and interpretation notes)
Start Your Project with MtoZ Biolabs
With extensive experience in RNA centric interaction profiling and functional genomics, MtoZ Biolabs delivers professional ChIRP-Seq Service.
Contact us to discuss your experimental design or request a quote. Our technical specialists are available to provide a free business assessment.
How to order?







