Bioinformatic Analysis Of Phosphorylated Proteomes
- Data sparsity and missing value handling: phosphopeptides often exhibit substantial missing values, requiring cautious application of imputation strategies (such as KNN, minimum value substitution, and Bayesian inference);
- Integration of site-level and protein-level analyses: effective mapping and integration between phosphorylation site information and protein-level interpretation must be achieved;
- Quantitative consistency and batch effects: experimental reproducibility and sample processing variability must be controlled through normalization methods (such as VSN and Z-score normalization).
Protein phosphorylation is one of the core regulatory mechanisms of cellular signal transduction and is widely involved in key biological processes such as cell growth, differentiation, apoptosis, metabolism, and disease progression. The acquisition of phosphoproteomic data through high-throughput mass spectrometry has become a routine practice in modern life science research; however, truly elucidating its biological significance relies on systematic and precise bioinformatic analysis of phosphorylated proteomes. Bioinformatic analysis of phosphorylated proteomes employs computational approaches for functional annotation, pathway enrichment analysis, and protein interaction network construction, thereby systematically revealing the regulatory mechanisms of phosphorylation events under different physiological or pathological conditions. Such analyses not only facilitate the identification of key kinase-substrate relationships and regulatory pathways but also enable the discovery of potential biomarkers and therapeutic targets. Furthermore, through time-series analysis, differential phosphorylation analysis, and subcellular localization prediction, researchers can gain deeper insight into the roles of protein phosphorylation in dynamic cellular processes. Therefore, bioinformatic analysis of phosphorylated proteomes represents a critical bridge between experimental data and functional interpretation and plays an indispensable role in both basic research and translational medicine.
Core Workflow of Bioinformatic Analysis of Phosphorylated Proteomes
1. Raw Data Processing and Quality Control
The first step in phosphoproteomic data analysis involves processing and identification of raw MS data, typically using software such as MaxQuant, Proteome Discoverer, and FragPipe:
(1) Peptide identification and quantification: database search algorithms (such as Andromeda and Sequest) are used to confirm peptide sequences and assign quantitative intensities.
(2) Phosphorylation site localization scoring (Site Localization): methods such as PhosphoRS and Ascore are applied to determine the precise position of phosphorylation sites.
(3) Evaluation of quality control metrics: including peptide coverage, proportion of phosphorylated peptides, peptide redundancy, and correlation among quality control samples.
2. Differential Analysis and Clustering
For multi-group comparative studies, differential analysis constitutes the core step in identifying biologically relevant changes:
(1) Statistical testing: commonly used methods such as t-tests, ANOVA, and Limma are applied to identify significantly differentially phosphorylated peptides.
(2) Multiple hypothesis correction: control of the false discovery rate (FDR) is implemented to reduce false positives.
(3) Clustering analysis: heatmaps, principal component analysis (PCA), and related approaches are used to reveal sample similarity and phosphorylation patterns.
3. Functional Annotation and Pathway Enrichment Analysis
After identifying differentially phosphorylated proteins, further exploration of their potential biological significance is required:
(1) GO enrichment analysis (Gene Ontology): to elucidate regulated molecular functions, cellular components, and biological processes.
(2) KEGG pathway analysis: to identify regulated signaling pathways, such as MAPK, PI3K-Akt, and mTOR.
(3) Protein-protein interaction networks (PPI): regulatory networks can be constructed using STRING or Cytoscape to identify key hub nodes.
4. Dynamic Phosphorylation Regulatory Analysis
Phosphorylation represents a dynamic and rapidly responsive regulatory mechanism. Researchers frequently employ:
(1) Time-series analysis (Time-series Clustering): to identify phosphorylation events that are activated or inhibited at specific time points.
(2) Kinase-substrate relationship prediction (Kinase-Substrate Analysis): tools such as Kinase Enrichment Analysis (KEA), iGPS, and NetworKIN are used to predict changes in upstream kinase activity.
(3) Motif analysis: to extract conserved amino acid sequence patterns surrounding specific phosphorylation sites and infer substrate specificity.
Common Challenges in Phosphorylation Data Analysis
Despite continuous technological advances, bioinformatic analysis of phosphorylated proteomes still faces multiple challenges:
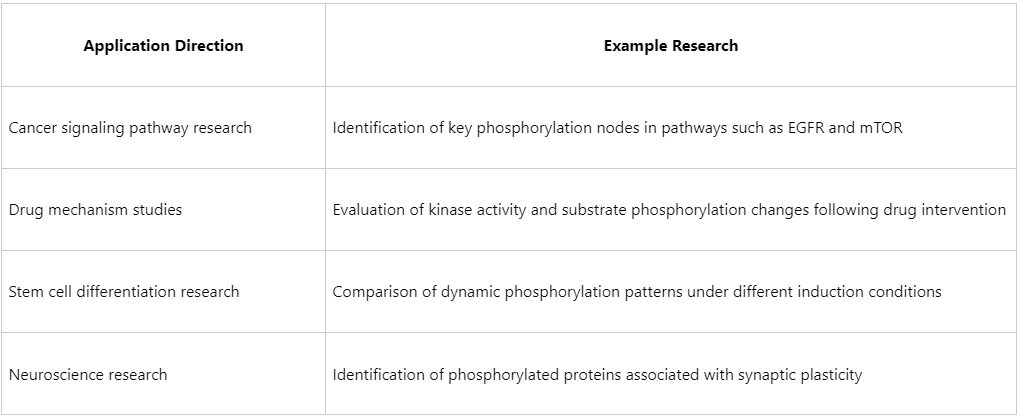
Application Scenarios: From Signaling Pathway Decoding to Clinical Translation
Bioinformatic analysis of phosphorylated proteomes is widely applied in both basic research and translational medicine:
Bioinformatic analysis of phosphorylated proteomes is rapidly emerging as a powerful approach for elucidating complex biological regulatory networks. With the continuous advancement of mass spectrometry technologies and iterative development of computational algorithms, the field is moving closer to dynamic, systematic, and precise decoding of cellular signaling. MtoZ Biolabs remains committed to translating advanced proteomic technologies into robust and practical solutions for researchers. Whether you are investigating cancer signaling pathways, identifying disease biomarkers, or developing targeted therapeutics, we are dedicated to serving as your trusted partner.
MtoZ Biolabs, an integrated chromatography and mass spectrometry (MS) services provider.
Related Services
How to order?







