Advantages and Disadvantages of High-throughput PPI Detection
-
Yeast Two-Hybrid (Y2H)
-
Affinity Purification coupled with Mass Spectrometry (AP-MS)
-
Biotin-based proximity labeling systems (BioID, TurboID, etc.)
-
Protein microarrays
-
Protein Interaction Chips (PPI Chips)
-
Computational prediction combined with experimental validation (e.g., in silico prediction + wet-lab verification)
-
Enables detection of direct binary interactions, facilitating analysis of domain-domain binding relationships.
-
Technically straightforward and well-established, suitable for large-scale interaction pair screening.
-
Relatively low cost, ideal for preliminary mapping of interaction networks.
-
Restricted to nuclear expression, unsuitable for membrane proteins or organelle-specific interactions.
-
Prone to false positives and negatives, necessitating further validation.
-
Does not capture dynamic interactions in native cellular environments.
-
Captures protein complexes under near-physiological conditions.
-
Supports construction of the full interactome.
-
Combined with high-resolution mass spectrometry, provides quantitative and structural information on interacting proteins.
-
Primarily detects stable complexes, less effective for transient or weak interactions.
-
Susceptible to non-specific binding, requiring appropriate controls (e.g., tagged protein background, empty vector).
-
Tagging or overexpression may introduce artificial artifacts.
-
Captures transient and weak interactions, suitable for dynamic interaction network studies.
-
Applicable to specific subcellular localizations or membrane protein interactions.
-
Flexible labeling and technically manageable.
-
Broad labeling can reduce specificity, requiring strict control of background signals.
-
Extensive biotinylation may complicate downstream mass spectrometry analysis.
-
Tagging may affect localization or function of the target protein.
-
In situ high-throughput detection, enabling thousands of protein interactions in a single experiment.
-
Amenable to standardization and automation, suitable for drug screening and antibody specificity assays.
-
Dependent on purified protein expression, folding and activity may be limited.
-
Predominantly in vitro, not fully reflecting intracellular interaction environments.
-
Not suitable for multi-protein complex detection.
-
High-density interaction detection using microfluidics or surface plasmon resonance platforms with real-time monitoring and automation.
-
Quantitative analysis of binding affinities (Kd) and kinetics is advantageous for distinguishing strong and weak interactions.
-
Reusable for multiple rounds, enhancing detection efficiency and sample utilization; suitable for large-scale validation or drug screening.
-
Requires high protein purity, activity, and stability; protein preparation remains a bottleneck.
-
High cost, more appropriate for high-quality validation following candidate interaction screening.
-
Most platforms are unsuitable for interactions dependent on cofactors or colocalization.
-
Rapidly screens potential interactions genome-wide based on protein structure, co-expression, evolutionary covariation, molecular docking, etc.
-
Complements experimental limitations in throughput and coverage, improving efficiency and targeting of candidate interactions.
-
Provides valuable preliminary insights for new species, uncharacterized proteins, or rare pathways.
-
Predictions depend on algorithm parameters and training datasets, with risk of false positives/negatives.
-
Requires systematic validation through AP-MS, Y2H, or PPI Chips to establish biological relevance.
-
Highly dependent on protein structural completeness and annotation quality, potentially constrained by database updates.
-
Rapidly construct high-quality interaction networks.
-
Identify potential targets and regulatory mechanisms.
-
Provide reliable experimental validation strategies.
Protein-Protein Interactions (PPIs) are central to biological processes such as intracellular signal transduction, metabolic regulation, and transcriptional control. With the rapid advancement of systems biology, the demand for high-throughput PPI detection techniques has grown significantly. These approaches not only elucidate dynamic interactions in complex biological systems but also facilitate drug target discovery, disease mechanism analysis, and functional annotation.
Currently, commonly employed high-throughput PPI detection techniques include:
Each technique has specific applications and inherent limitations. The following provides a systematic comparison based on technical principles, throughput, sensitivity, specificity, application scenarios, and scalability.
Overview of Common High-throughput PPI Detection Techniques
1. Yeast Two-Hybrid (Y2H)
(1) Advantages
(2) Limitations
2. Affinity Purification-Mass Spectrometry (AP-MS)
(1) Advantages
(2) Limitations
3. Biotin-based Proximity Labeling (BioID / TurboID / APEX, etc.)
(1) Advantages
(2) Limitations
4. Protein Microarrays
(1) Advantages
(2) Limitations
5. Protein Interaction Chips (PPI Chips)
(1) Advantages
(2) Limitations
6. Computational Prediction with Experimental Validation (In silico + Wet-lab)
(1) Advantages
(2) Limitations
Technical Comparison and Recommended Applications
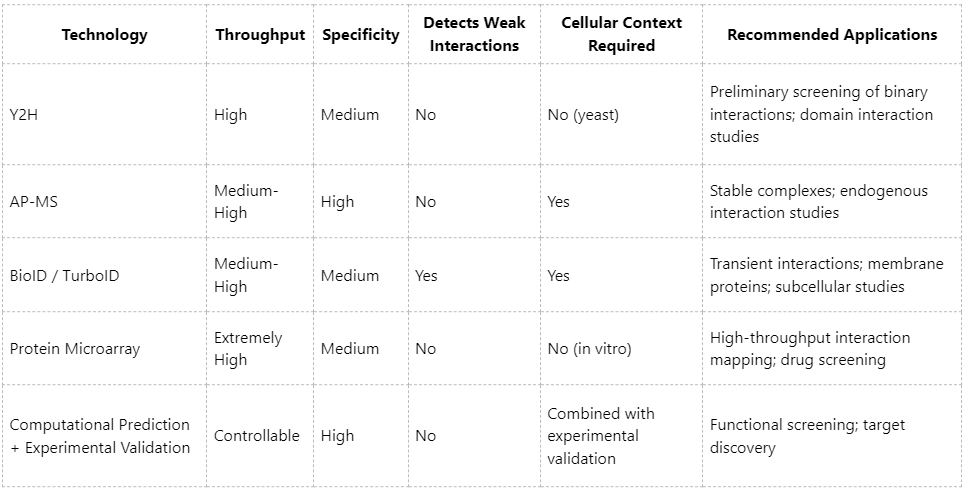
No single high-throughput PPI technique can fully capture all types of protein interactions. Therefore, integrating multiple approaches is the current standard. For instance, Y2H may be used for initial candidate screening, followed by AP-MS or BioID for validation in cellular contexts. Integration with proteomics, metabolomics, and functional assays can further enhance the depth and reliability of PPI network analyses.
At MtoZ Biolabs, we offer end-to-end PPI detection services including AP-MS, BioID, TurboID, and comprehensive data analysis. Leveraging high-resolution mass spectrometry and expert bioinformatics support, we help researchers:
Contact us to design customized interactomics solutions and accelerate your research discoveries.
MtoZ Biolabs, an integrated chromatography and mass spectrometry (MS) services provider.
Related Services
How to order?







