4D Methylation Proteomics Service
- Provide sample group definitions, replicate structure, and study goals
- Keep samples cold, minimize freeze-thaw cycles, and ship under appropriate cold conditions
- Document lysis buffer components and any inhibitors used
- Avoid introducing polymer contaminants and excessive salts when possible
- Raw LC-MS/MS files (.raw)
- Processed data tables (CSV or Excel format) containing identified and quantified protein methylation
- Summary reports in PDF format, including analytical methods, quality control results, and key findings
- Figures and chromatograms in high-resolution image formats (TIFF or PNG)
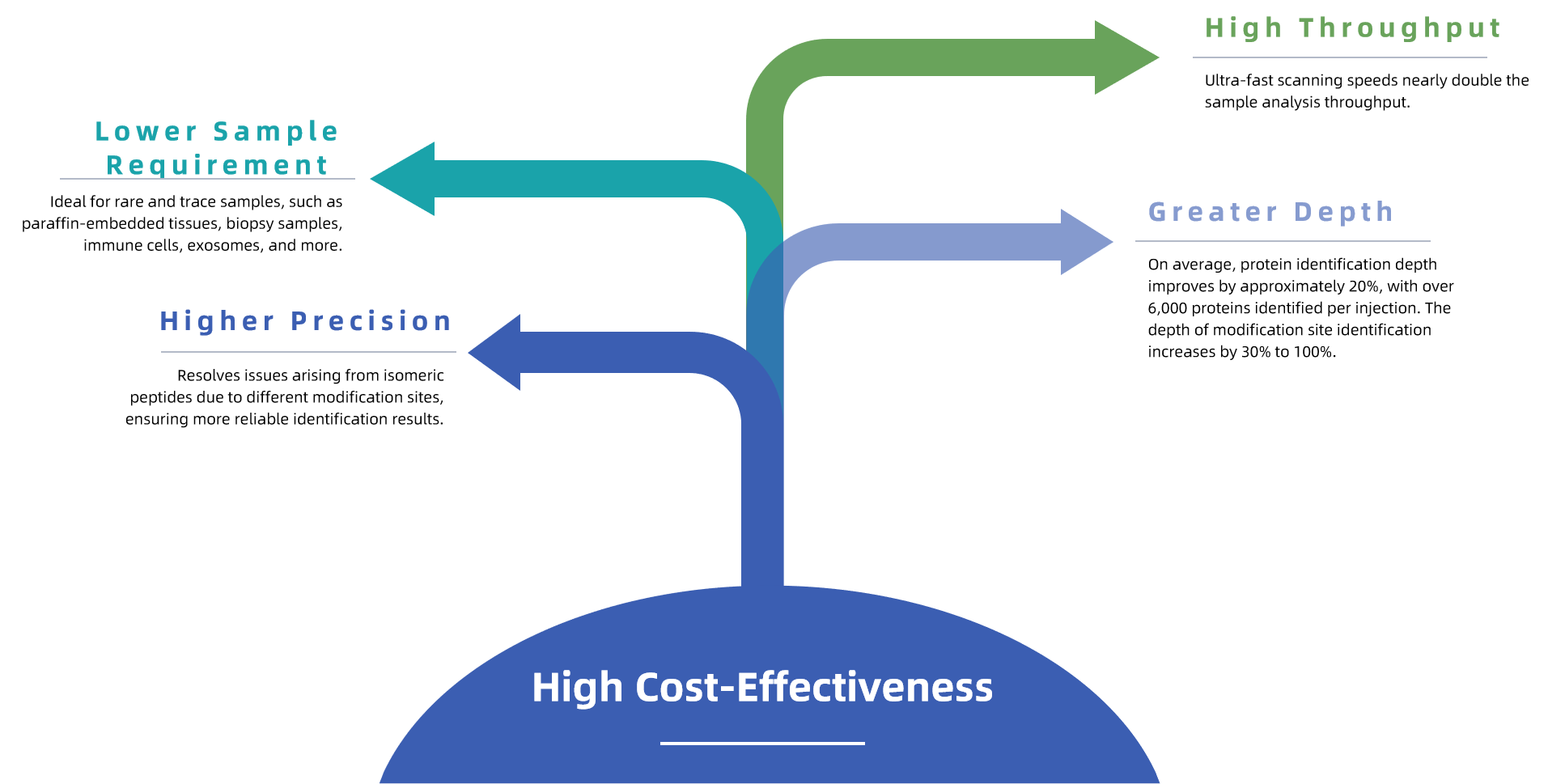
MtoZ Biolabs offers 4D Methylation Proteomics Service to deliver high-quality insights into protein methylation across complex biological samples. This service combines ion mobility spectrometry with mass spectrometry (MS) to enhance the detection, sensitivity, and throughput of methylation site identification, making it ideal for methylome profiling, mechanism studies, and biomarker discovery.
Technical Principles
Protein methylation most commonly occurs on lysine and arginine residues and can appear in multiple methylation states. Proteome-wide detection typically benefits from enrichment because methylated peptides are often low-abundance and can be masked by the background peptide pool. Immunoaffinity enrichment uses methylation-specific binders to selectively capture methylated peptides prior to LC-MS/MS, improving depth and site-level confidence.
4D proteomics adds ion mobility separation as an additional dimension beyond retention time, m/z, and intensity. On the timsTOF Pro platform, trapped ion mobility spectrometry separates ions by mobility, which reflects molecular shape and collisional cross section, before MS/MS acquisition. This additional separation reduces spectral interference and improves precursor selectivity in complex samples. Combined with fast acquisition strategies, the 4D workflow increases identification depth and sensitivity while helping reduce sample consumption and overall scan time for methylation proteomics projects.
4D Methylation Proteomics Service at MtoZ Biolabs
MtoZ Biolabs uses the timsTOF Pro mass spectrometer, integrated with ion mobility spectrometry and PASEF, to offer a robust 4D methylation proteomics workflow. This technology supports methyl-peptide enrichment for capturing methylated peptides, followed by high-resolution analysis that provides deeper and more accurate profiling of methylation sites. Our service includes comprehensive reporting of methylation site identification, site localization, and quantitative analysis.
Sample Submission Guidelines
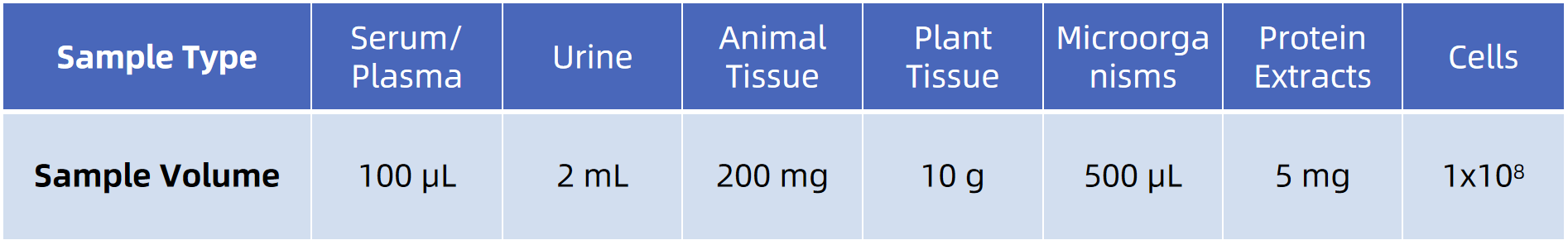
To support methylation preservation and reliable quantification, submit samples with consistent handling across groups.
For more information, please refer to Sample Submission Guidelines for Proteomics. If you are unsure about sample preparation, our team is available for consultation and can guide you through the process to ensure optimal results.
Applications of 4D Methylation Proteomics Service
● Profiling methylation dynamics in response to treatments or disease models
● Characterizing methylation in chromatin-associated proteins, transcriptional regulators, and RNA-binding proteins
● High-throughput methylation analysis for biomarker discovery and drug target identification
● Investigating methylation in complex biological samples such as tissues, biofluids, and cell lysates
Start Your Project with MtoZ Biolabs
If you need high-confidence methylation site detection with the enhanced sensitivity of ion mobility spectrometry, contact MtoZ Biolabs today. Our expert team will work with you to design the best workflow based on your research goals and sample type.
FAQ
Q1: What is the service general workflow?
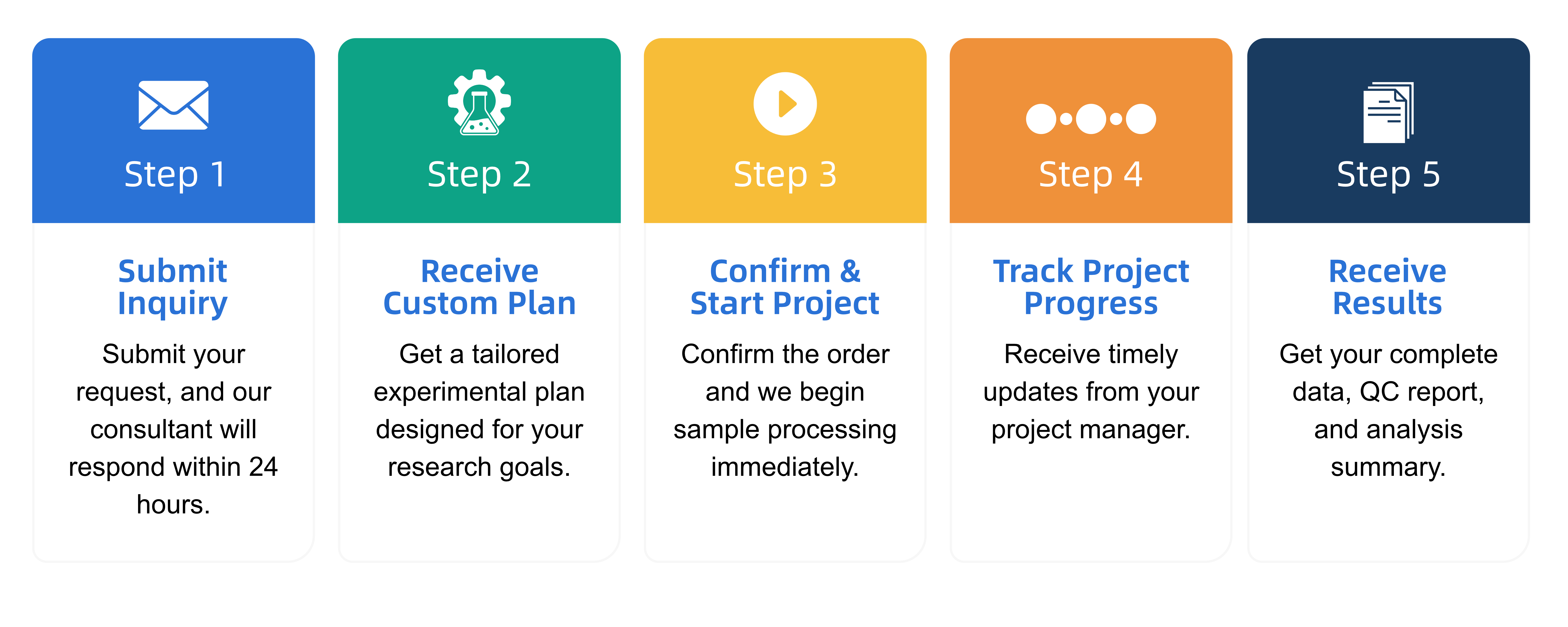
Q2: What data formats are provided?
MtoZ Biolabs provides results in multiple standard formats to ensure compatibility with various analysis and visualization tools. Deliverables typically include:
Additional file formats or data structures can be provided upon request to meet specific research or publication requirements.
Related Services
How to order?







