Spatial Metabolomics Service
- Cost-effective and label-free in situ detection
- Eliminates the need for extraction, thereby reducing human error
- Offers high spatial and mass resolution along with exceptional sensitivity
- Features a wide detection range, allowing for both targeted analysis of specific metabolites and broad screening
- Animal tissues: Including organs from mice and rats such as brain, kidney, heart, liver, spleen, spinal cord, pituitary gland, pancreas, and skin.
- Cancer samples: From cancers like esophageal, thyroid, lung, breast, stomach, glioma, nasopharyngeal, skin, and liver.
- Plant tissues: Focusing on roots and seeds.
- Cancer Research: Examining metabolic alterations in tumor tissues to identify potential biomarkers and elucidate their biological roles.
- Drug Development: Evaluating the effects of pharmacological compounds on metabolic networks to enhance the understanding of drug mechanisms.
- Plant Metabolism Research: Investigating the metabolic responses of plants under various environmental conditions to uncover adaptive strategies.
- Neuroscience: Analyzing metabolic discrepancies in neural tissues to contribute to research on neurodegenerative disorders.
- Comprehensive Experimental Details
- Materials, Instruments, and Methods
- Total Ion Chromatogram & Quality Control Assessment (project-dependent)
- Data Analysis, Preprocessing, and Estimation (project-dependent)
- Bioinformatics Analysis
- Raw Data Files
Spatial metabolomics is a state-of-the-art molecular imaging technology that integrates mass spectrometry analysis with imaging visualization. This label-free technique facilitates the in situ characterization of metabolites within biological samples, effectively addressing the qualitative, quantitative, and spatial distribution challenges associated with these metabolites. Its applications are extensive in both physiological and pathological contexts, particularly in cancer research, drug development, and environmental monitoring. At MtoZ Biolabs, we are dedicated to delivering high-quality spatial metabolomics analysis services to support comprehensive biological investigations.
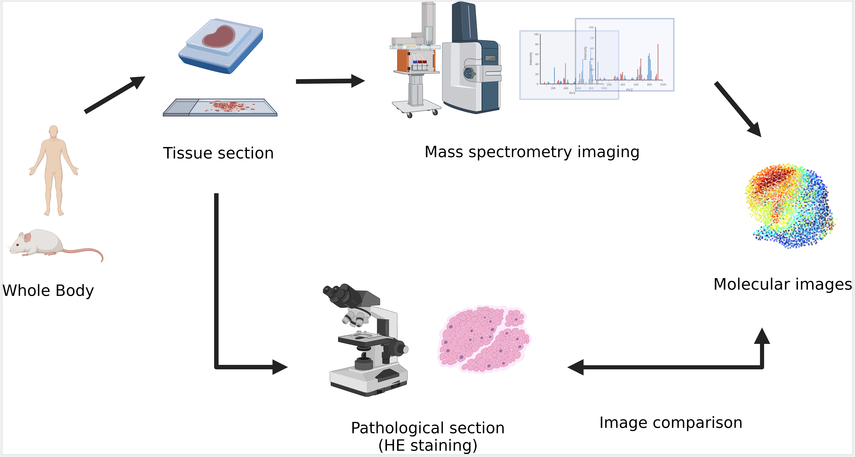
Zheng, Y. et al. Front Oncol. 2023.
Figure 1. The Process of Transformation in Multi-Omics in the Research of Human Tumor’s Biology
Technical Principles
Matrix assisted laser desorption ionization-mass spectrometry (MALDI-MS) operates on the principle that laser irradiation of a matrix-analyte mixture generates a co-crystal. The energy absorbed by the matrix molecules leads to thermal expansion, resulting in the sublimation of the matrix crystals. During the detection phase, the sample surface is divided into sections using a grating, allowing for the collection of spectra point-by-point. This process determines the mass-to-charge ratios and ion intensities from each pixel on the sample surface. When integrated with mass spectrometry imaging software, this approach creates a two-dimensional distribution map of the corresponding molecules or ions on the sample surface.
Service Advantages
Why Choose MtoZ Biolabs?
Sample Submission Requirements
Applications
Deliverables
What Could be Included in the Report?
MtoZ Biolabs is committed to partnering with you to tailor solutions that meet your project's unique demands. For further inquiries or specific needs related to spatial metabolomics, please contact us. Our team of experts is ready to provide specialized advice and support.
How to order?







