Small Molecule Actives Identification and Quantification
Small molecule drugs, predominantly chemically synthesized drugs with molecular weight lower than 1000 Da. These drugs are characterized by their widespread use and mature theoretical foundations. Statistically, small molecule drugs comprise about 98% of commonly used medications. They have excellent dispersibility and druggability due to their structural and chemical properties. These properties grant small molecule drugs significant advantages in drug research and development as well as in various pharmaceutical domains. Another notable feature of small molecule drugs is they are easy to obtain by chemical synthesis, which facilitates modifications in their chemical structure and improvements in their physical and chemical properties.
The development of small molecule drugs traces back to 1828, marked by the successful synthesis of urea from inorganic reagents and the transformation of carbon disulfide into acetic acid. By the end of the 19th century, the chemical industry had produced a series of chemical dyes and other chemicals, leading to the launch of aspirin (acetylsalicylic acid, Bayer, 1899), a blockbuster in the pharmaceutical industry. Seventy years later, with the synthesis of Vitamin B12 by Woodward and Eschenmoser, the field of synthetic organic chemistry reached maturity. The era of sensational advancements in organic synthesis dawned with pharmaceutical companies and research institutions developing a plethora of small molecule drugs, thereby offering new options for treating diseases and extending life span.
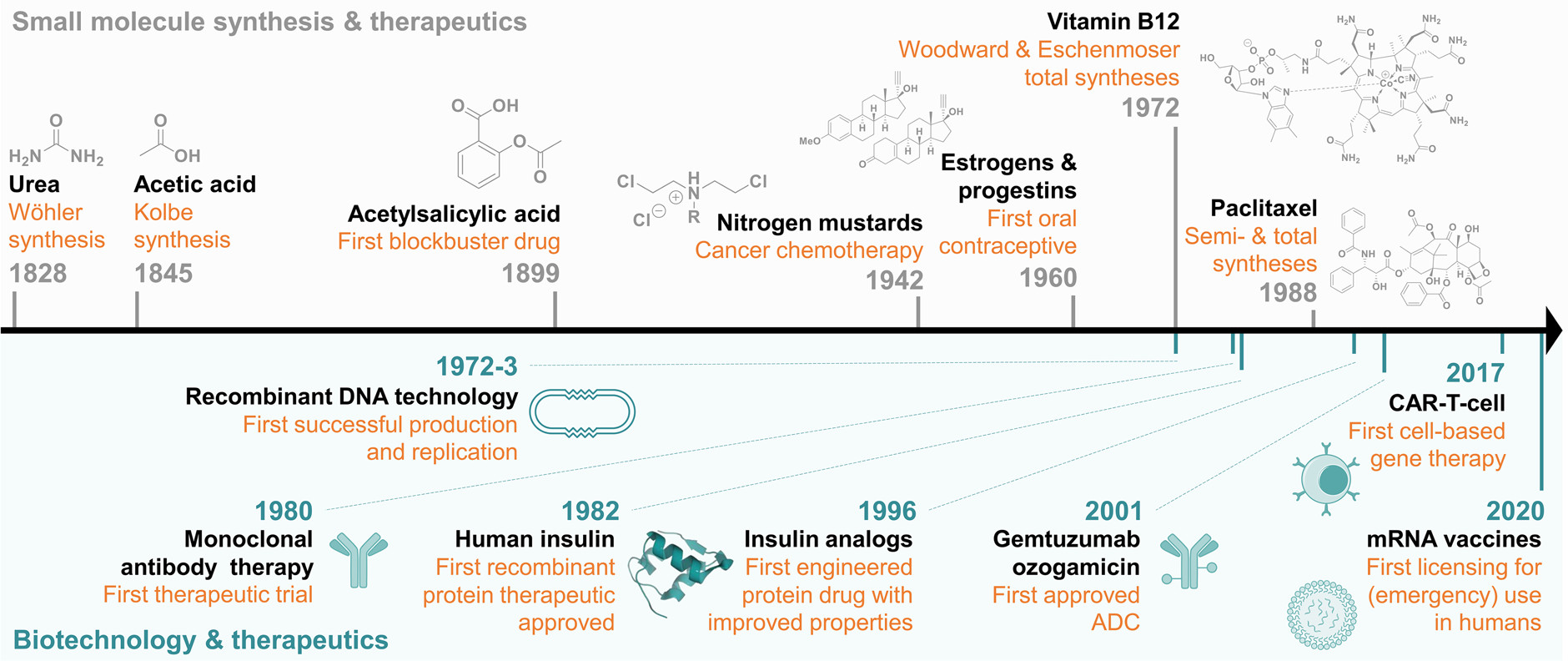
Figure 1. The Development of Small Molecule Drugs[1]
A variety of small molecule inhibitors are used to treat cancer. Typically functioning as signal transduction inhibitors, these drugs specifically block signaling pathways essential for tumor growth and proliferation to cure diseases. Although most of these approvals are for extending the survival of patients with hard-to-treat, advanced-stage cancers traditionally treated with chemotherapy. Many of these drugs have shown superiority over cytotoxic chemotherapy, having fewer side effects as first-line treatments in relapsed or metastatic cases. Some small molecule inhibitors are also approved for treating minimal residual disease or as adjuvant therapies.
Small molecule drugs commonly exert their effects by interacting with receptors located on cell surfaces, within the nucleus, or in the cytoplasm. Receptors are large molecules responsible for activating chemical signals within or between cells. Molecules that bind to receptors are known as ligands. Ligands can act as full agonists, partial agonists, inverse agonists, and antagonists, depending on their affinity and efficacy, which are determined by their chemical structure. Most approved small molecule inhibitors target intracellular kinases, regulating cellular signaling by transferring phosphate groups to target proteins. Currently, a wide range of targets is being explored, including those involved in protein-protein interactions, cancer metabolism, and immune regulation.
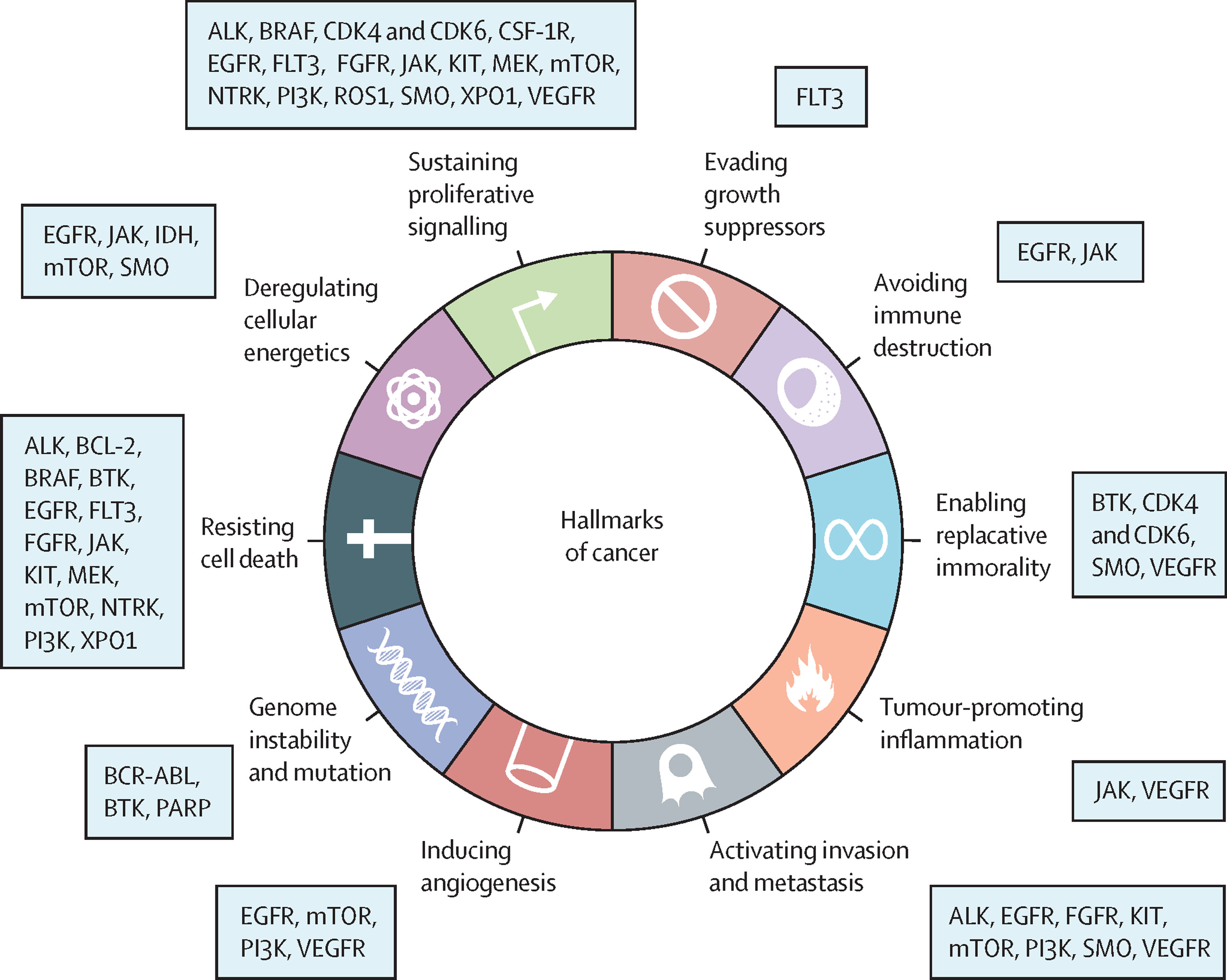
Figure 2. Small Molecule Inhibitors for Cancer Treatment [2]
Small molecules play a crucial role as chemical probes in biomedical research, aiding in the understanding the development of disease. Traditional small molecule drugs have a priority in drug research. Today, novel biological approaches like proteolysis-targeting chimeras (PROTACs) and RNA-targeting small molecules (RSMs) have broadened the research field for small molecule drugs.
The development and analysis of small molecule drugs encompass the process of their medicinal chemistry, metabolite identification, impurity profiling, drug metabolism, and pharmacokinetics. The successful testing and release of small molecule drugs rely on efficient analytical tools, validation guidelines, and standards. Various methods can achieve the separation and identification of small molecules. They include wet chemistry, high-performance liquid chromatography (HPLC), gas chromatography (GC), thin-layer chromatography (TLC), capillary electrophoresis (CE), nuclear magnetic resonance spectroscopy (NMR), and mass spectrometry (MS).
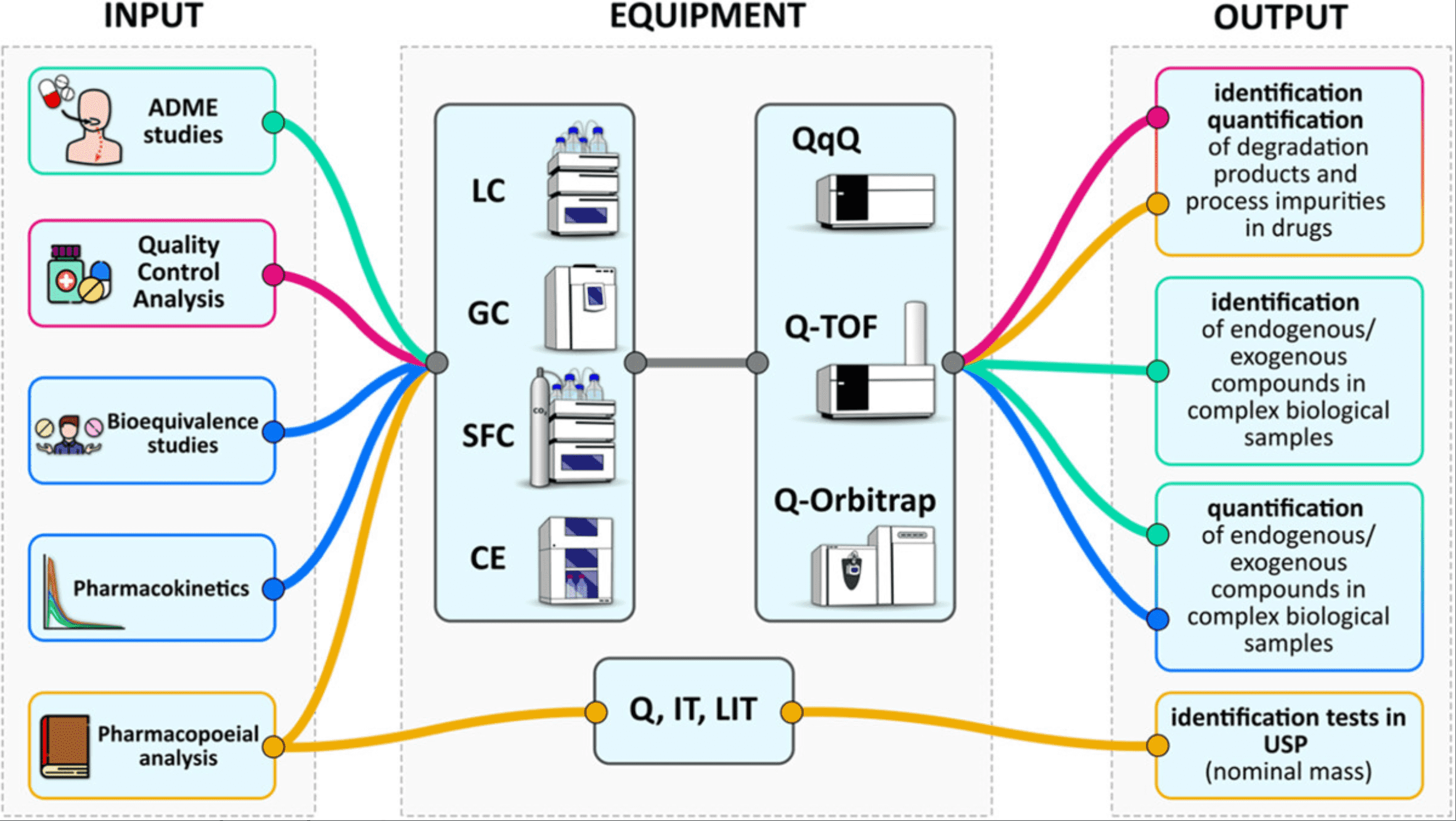
Figure 3. The Application of MS in the Pharmaceutical Analysis of Small Molecule Chemical Drugs [3]
The analysis of small molecules based on HPLC typically employ reverse-phase separation techniques. Hydrophilic interaction chromatography (HILIC) and normal-phase chromatography both can be applied into the separation of polar compounds, with the former often yielding more precise results. Ionizable compounds can be separated using ion exchange methods, while organic anions or cations may also be separated through ion chromatography. HPLC columns can be packed with fully porous silica particles, surface porous silica particles, polymer particles, or monolithic silica rods as the stationary phase, and may also be packed with alumina, zirconia, and carbon particles. The pore size of the stationary phase material typically ranges from 60Å to 160Å. The particle size for HPLC stationary phases generally ranges from 3µm to 5µm, while particle sizes for ultra performance liquid chromatography (UHPLC) are smaller than 2µm. Column selectivity can be varied through the modification of the column. The modification of C18 alkyl chain is the most commonly used chemical modification for reverse-phase chromatography (RPC). However, a wide variety of polar modifications for stationary phases, as well as modifications offering ion-exchange or chiral properties, allow for the separation of nearly all compounds soluble in liquid, through modifications like C30, C8, phenyl, and pentafluorophenyl. The mobile phase in RP-HPLC typically contains aqueous buffer solutions or water and water-miscible organic solvents, such as acetonitrile or methanol.
MS is a critical technique for the analysis of small molecule drugs characterized by high sensitivity, a broad range of molecular mass analysis, and abundant molecular structure information. The liquid chromatography coupled to tandem mass spectrometry (LC-MS/MS) method combines highly efficient separation of LC with the high sensitivity of MS, providing a two-dimensional separation based on the mass-to-charge ratio (m/z). This is essential for potential candidate drugs inpharmacokinetics and pharmacodynamics researches and preclinical and clinical studies. Liquid chromatography-MS (LC-MS) methods ensure the quality and safety of drugs, including the identification and quantification of impurities and degradation products.
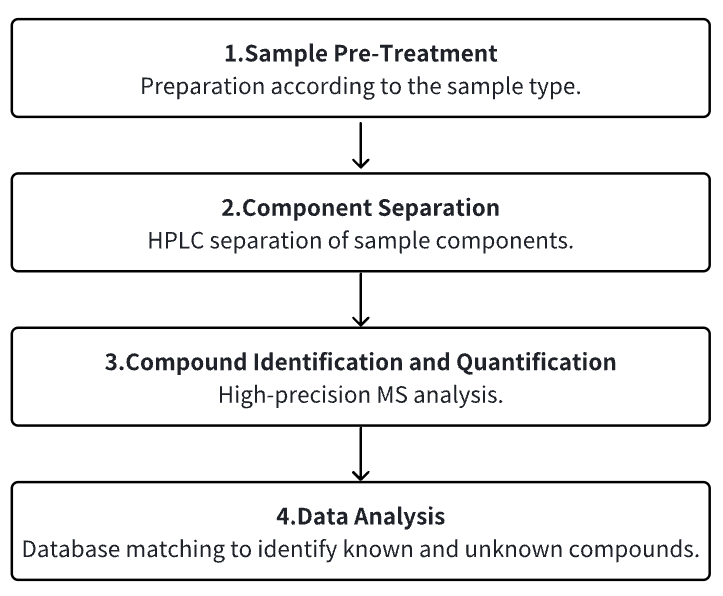
Analysis Workflow
1. Sample Pre-Treatment
2. Component Separation
3. Compound Identification and Quantification
4. Data Analysis
Service Advantages
1. Accurate identification and quantification of compounds across various sample types.
2. High-reliability, high-precision MS analysis.
3. Comprehensive bioinformatics analysis.
Sample Results
1. Study of Pharmacokinetics and Metabolomic Profiles of Novel Potential PLK-1 Inhibitors Using UHPLC-MS/MS and UHPLC-Q-Orbitrap/HRMS [4]
Polo-like kinase 1 (PLK-1) plays a crucial role in cell division, and its aberrant expression is closely related to various cancers. It has been targeted for the development and study of small molecule inhibitors in clinical trials. In this study, we designed and synthesized a potent new PLK-1 inhibitor, compound 7a. We have established and validated a UHPLC-MS/MS method for the exploration of the pharmacokinetic properties of this drug candidate. This method is characterized by excellent selectivity, high sensitivity, and a broad linear range.
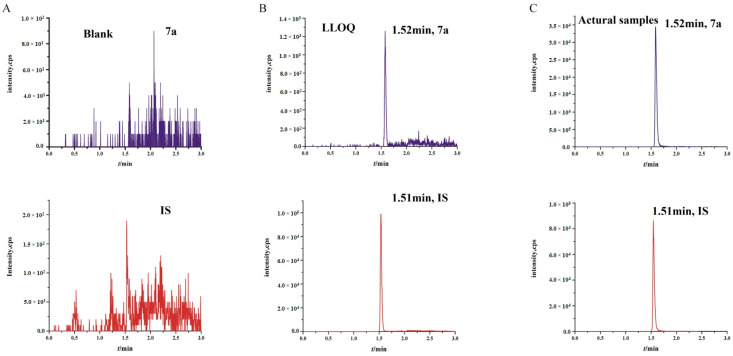
Figure 4. Representative Chromatogram of Drugs in Rat Plasma
2. Ultra-sensitive Immunoaffinity 2D-LC-MS/MS Method for Evaluating Covalent Inhibitor Target Engagement of KRAS G12C in Tumor Biopsies [5]
KRAS is one of the most frequently mutated oncogenes, with KRAS G12C recently emerging as a tractable target for small molecule intervention. GDC-6036, an investigational KRAS G12C inhibitor, functions by irreversibly binding to the switch II pocket of KRAS G12C in its inactive GDP-bound state, thereby inhibiting GTP binding and activation. Assessing target engagement is crucial for clinical drug development, demonstration of mechanism of action, guidance on dosage usage, understanding of its pharmacodynamics, and exploration of drug resistance. This study reports the development of an ultra-sensitive method for assessing KRAS G12C engagement. Through the combination of immunoaffinity enrichment using commercial anti-RAS antibodies and targeted 2D-LC-MS/MS technology, we achieve quantitative measurement of both free and GDC-6036-bound KRAS G12C proteins. This assay was applied in a KRAS G12C-positive non-small cell lung cancer xenograft model treated with GDC-12 to evaluate the feasibility of analyzing fine-needle biopsies.
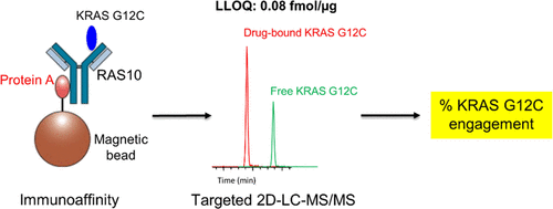
Figure 5. Evaluation of Immunoaffinity Involving KRAS G12C Using 2D-LC–MS/MS
3. Development and Validation of a Sensitive and Specific LC-MS/MS Method for Wnt Signaling Inhibitor IWR-1-Endo: Application in Brain Microdialysis Studies [6]
IWR-1-endo, a small molecule inhibitor, effectively blocks the Wnt/β-catenin signaling pathway by stabilizing the AXIN2 destruction complex, potentially inhibiting drug efflux at the blood-brain barrier. In brain microdialysis studies on mice, we established a validated, sensitive, and robust LC-MS/MS method to determine the concentrations of IWR-1-endo in mouse plasma and brain microdialysis fluids. The LC-MS/MS method was successfully applied in mouse pharmacokinetics and brain microdialysis studies to characterize the exposure of unbound IWR-1-endo in brain extracellular fluid and plasma.
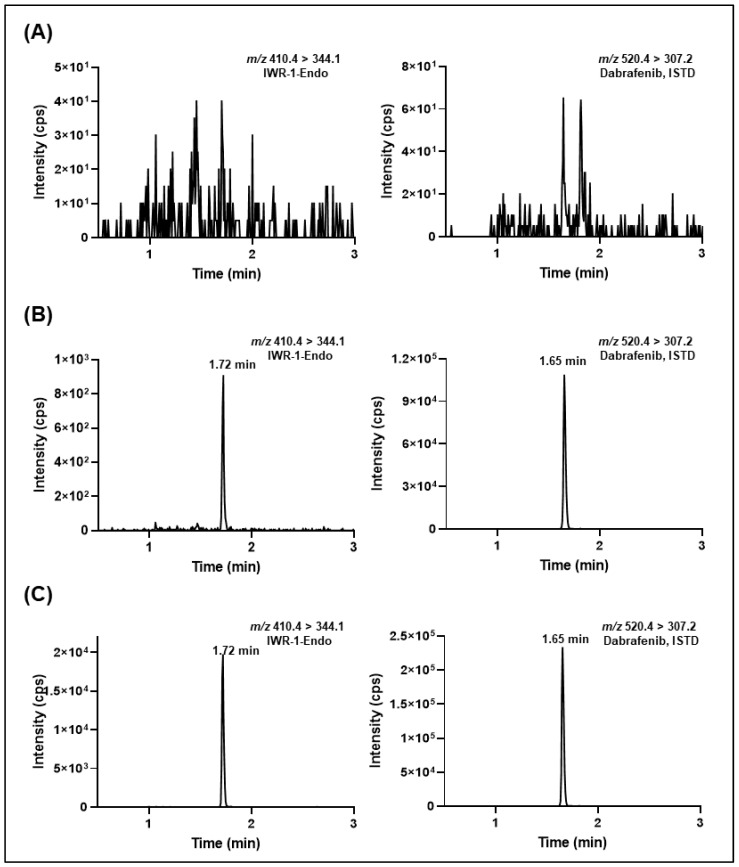
Figure 6. Representative Chromatogram of Dialysates from Various Groups
4. Structural MS Detection of Conformational Activation and Dissociation Induced by Inhibitors in CDK12/CDK13-Cyclin K Complex [7]
The integration of lysine-reactivity profiling-MS (LRP-MS) with native MS (nMS) as structural MS approaches unveiled a novel inhibition mechanism where small molecule inhibitor SR-4835 induced conformational activation and dissociation of the cell cycle kinase 12/13-Cyclin K complex (CDK12/CDK13-Cyclin K), paving new avenues for the rational design of CDK12/CDK13 small molecule inhibitors.
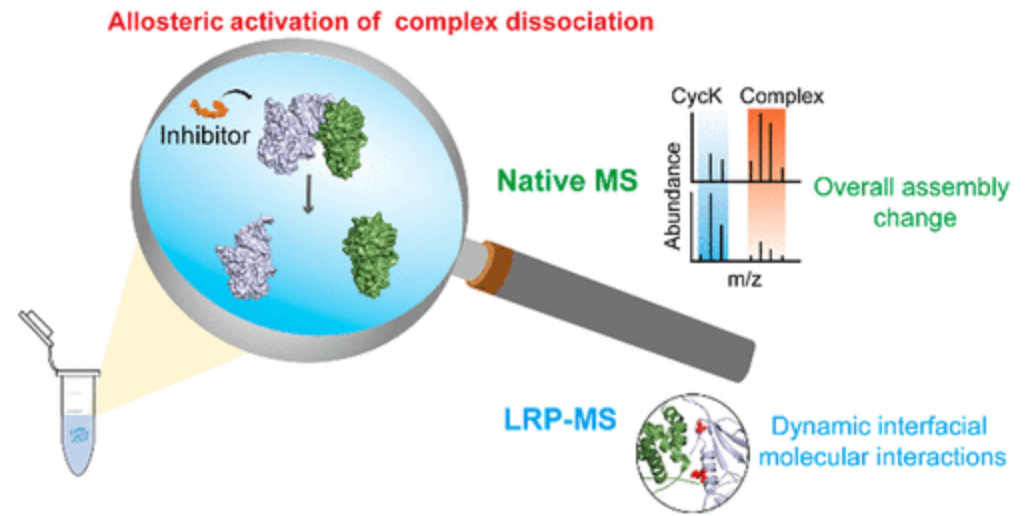
Figure 7. Structural MS Analysis of Conformational Activation Induced by Inhibitor Dissociation of CDK12/CDK13-Cyclin K
Sample Submission Requirements
1. Kindly submit samples that are as highly purified as feasible.
2. Endeavor to minimize contamination with impurities.
Services at MtoZ Biolabs
1. Comprehensive Experimental Procedure
2. Relevant Instrument Parameters
3. Raw MS Data
4. Detailed Report on the Identification of Small Molecular Compounds
Applications
1. Drug Metabolite Analysis
Drug metabolites arise from the process of drug metabolism, whose structures closely depend on the type of metabolic reaction and the structural attributes of the precursor compounds. Upon metabolic transformation in the body, drugs generally undergo partial structural modifications without significant structural changes to the parent drug. Therefore, metabolites often share similar mass spectral characteristic ions with the parent drug, facilitating their identification. Combined with other fragment characteristics, this information allows for structural inference of metabolites. In analyzing drug metabolites, it's essential to compare the electron impact MS spectra of metabolites with that of the synthetic products, with the chemical structure ultimately confirmed by NMR. Multi-stage MS (MS^n) can directly provide structural information about the parent molecule, eliminating the need for pure metabolite preparation, while biological samples only require simple pre-treatment.
2. Drug Target Identification
Proteomics based on MS facilitates the direct analysis of interactions between small molecules and the protein. Chemical proteomics can detect direct interactions between compounds and proteins, typically relying on chemical probes derived from bioactive compounds to enrich drug targets from cell extracts or live cell systems. Additionally, emerging methods exploit the effects of compound binding on protein structure and biophysical properties, such as thermal stability or resistance to proteolysis. Target identification of bioactive compounds usually relies on affinity enrichment, where bioactive molecules are derivatized with affinity tags (e.g., biotin) or bioorthogonal reactive groups for immobilization. Affinity enrichment allows for the isolation of proteins interacting with the functional probe during incubation with cell proteins.
3. Quality Control of Small Molecule Drugs
The structural characteristics, physicochemical properties, and quality control of small molecule drugs require the establishment of related quality control criteria. The content should conclude the impact of starting materials and reagents used in the preparation process, intermediates and by-products, and organic solvents on the final product quality. It is necessary to confirm the chemical structure and components of small molecule drugs and linker-small molecule drugs and conduct corresponding quality studies. Structural confirmation studies may include infrared, ultraviolet, nuclear magnetic resonance (carbon, hydrogen, and, when necessary, two-dimensional correlation spectra), and MS analyzes. Quality studies should cover aspects such as appearance, identification, examination, and content determination, with particular attention to coupled impurities and chiral isomers, imposing limits on unknown impurities. For small chiral molecule drugs, quality control of the starting materials and synthetic intermediates should be conducted, along with chiral isomer analysis of the final product as needed.
FAQ
Q1:How are small molecule compounds screened?
Screening of lead compounds can be efficiently conducted on a drug activity screening platform, which facilitates high-throughput screening, high-content screening, preliminary biological activity assays, and optimization. A synergistic approach combines physical compound libraries with virtual screening. Utilizing tools such as nanoliter-scale acoustic liquid handling systems, multimodal microplate detection systems, high-content confocal fluorescence microscopy, surface plasmon resonance interaction systems, and isothermal titration calorimetry (ITC) interaction systems, pharmacological activity screening is conducted with high efficiency.
References
[1] Roman BI. The Expanding Role of Chemistry in Optimizing Proteins for Human Health Applications. J Med Chem. 2021 Jun 10;64(11):7179-7188. doi: 10.1021/acs.jmedchem.1c00294. Epub 2021 May 20. PMID: 34014084.
[2] Bedard PL, Hyman DM, Davids MS, Siu LL. Small molecules, big impact: 20 years of targeted therapy in oncology. Lancet. 2020 Mar 28;395(10229):1078-1088. doi: 10.1016/S0140-6736(20)30164-1. PMID: 32222192.
[3] Khalikova M, Jireš J, Horáček O, Douša M, Kučera R, Nováková L. What is the role of current mass spectrometry in pharmaceutical analysis? Mass Spectrom Rev. 2023 Jul 28. doi: 10.1002/mas.21858. Epub ahead of print. PMID: 37503656.
[4] Wang L, Lei H, Lu J, Wang W, Liu C, Wang Y, Yang Y, Tian J, Zhang J. Study on Pharmacokinetics and Metabolic Profiles of Novel Potential PLK-1 Inhibitors by UHPLC-MS/MS Combined with UHPLC-Q-Orbitrap/HRMS. Molecules. 2023 Mar 10;28(6):2550. doi: 10.3390/molecules28062550. PMID: 36985522; PMCID: PMC10053003.
[5] Meng L, Chan EW, Ng C, Aimi J, Tran JC, Oh AJ, Merchant M, Purkey HE, Heffron TP, Kaur S, Xu K, Shi Z, He J. Assessment of KRAS G12C Target Engagement by a Covalent Inhibitor in Tumor Biopsies Using an Ultra-Sensitive Immunoaffinity 2D-LC-MS/MS Approach. Anal Chem. 2022 Sep 20;94(37):12927-12933. doi: 10.1021/acs.analchem.2c03146. Epub 2022 Sep 9. PMID: 36083155.
[6] Nair S, Davis A, Campagne O, Schuetz JD, Stewart CF. Development and Validation of a Sensitive and Specific LC-MS/MS Method for IWR-1-Endo, a Wnt Signaling Inhibitor: Application to a Cerebral Microdialysis Study. Molecules. 2022 Aug 25;27(17):5448. doi: 10.3390/molecules27175448. PMID: 36080214; PMCID: PMC9457781.
[7] Bai Y, Liu Z, Li Y, Zhao H, Lai C, Zhao S, Chen K, Luo C, Yang X, Wang F. Structural Mass Spectrometry Probes the Inhibitor-Induced Allosteric Activation of CDK12/CDK13-Cyclin K Dissociation. J Am Chem Soc. 2023 May 31;145(21):11477-11481. doi: 10.1021/jacs.3c01697. Epub 2023 May 19. PMID: 37207290.
How to order?







