Resources
Proteomics Databases
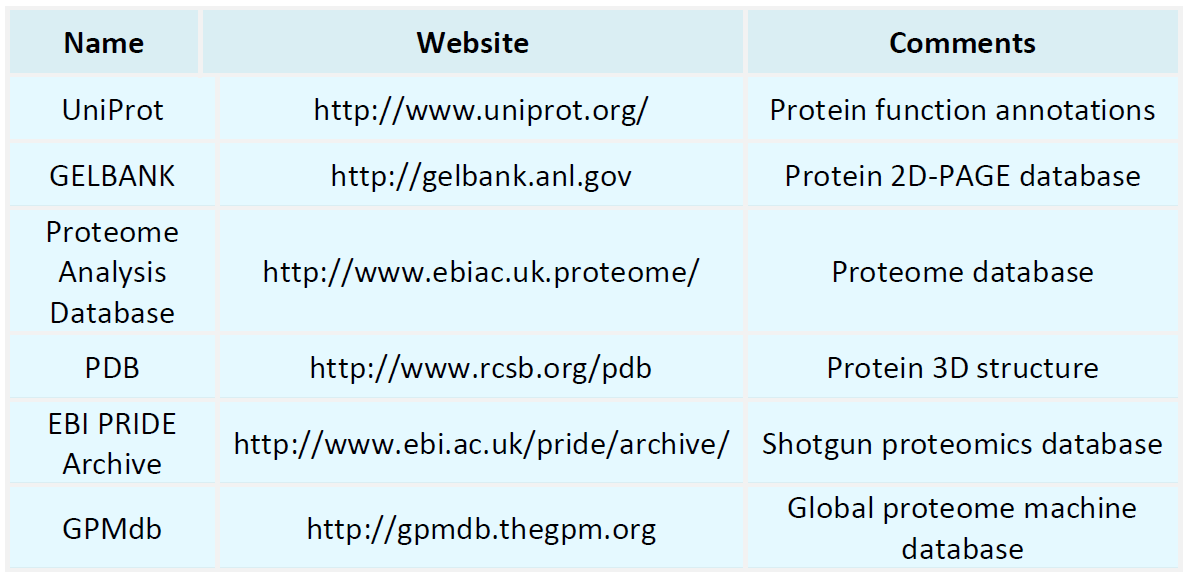
Metabolomics Databases
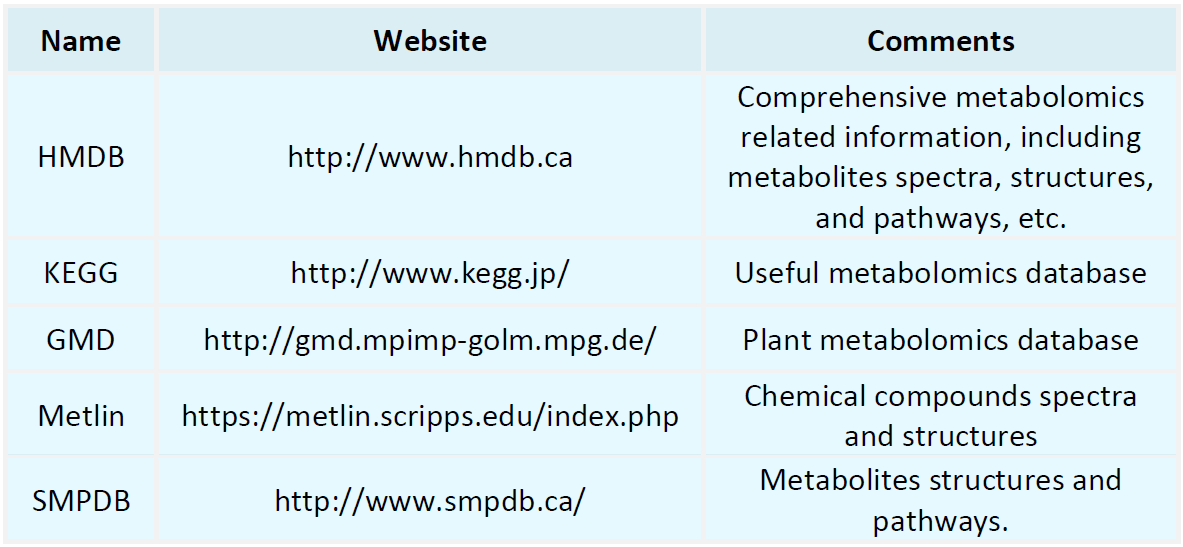
-
• Protein N-Terminal Sequencing Analysis Using MALDI-TOF Mass Spectrometry
Protein N-terminal sequencing is a critical approach for elucidating protein function, post-translational modifications, and biosynthetic pathways. The Edman degradation method, while a classical technique for N-terminal sequencing, is limited in its applicability to complex samples. In recent years, matrix-assisted laser desorption/ionization time-of-flight mass spectrometry (MALDI-TOF-MS) has emerged as a valuable tool in N-terminal sequencing analysis due to its high sensitivity, rapid analysis .........
-
• Overcoming Challenges in C-terminal Sequencing: A Practical Guide for Experimental Optimization
C-terminal sequencing is of significant value in proteomics research; however, experimental implementation often encounters considerable challenges due to the complex chemical properties of protein C-termini, limited enzymatic specificity, and difficulties in enriching C-terminal sequences. Optimizing experimental strategies to enhance sequencing sensitivity, specificity, and accuracy has become a central concern for researchers. In this guide, we explore optimization approaches for each critical .........
-
• Protein N-Terminal: A Comprehensive Analysis from Biological Functions to Sequencing Technologies
The protein N-terminal represents a critical element of both its structural integrity and functional capacity, directly influencing post-translational modifications (PTMs), subcellular localization, protein–protein interactions, and degradation pathways. Beyond serving as the physical marker for the initiation of protein synthesis, the N-terminal also functions as a central regulatory element governing cellular fate. However, the highly heterogeneous nature of its modifications and its complex biological...
-
• Protein C-Terminal Sequencing: Methods, Challenges, and Optimization
Protein serves as a central executor of biological functions, with its structure intricately linked to its function. While extensive efforts have been devoted to the sequencing of protein N-termini, studies on the C-terminus (carboxyl terminus) have comparatively lagged behind. In reality, the C-terminal region plays critical roles in regulating protein stability, subcellular localization, signal transduction, and degradation. Notably, C-terminal information has demonstrated distinct value in areas such....
-
• Efficient C-Terminal Sequencing Strategies: Enhancing Accuracy and Sensitivity
The C-terminus of proteins plays a critical role not only in determining their intracellular localization, stability, and biological activity, but also in mediating a variety of essential biological processes, including ubiquitin-dependent degradation, regulation of signal transduction, and proteolytic activation. Compared to the N-terminus, the C-terminal region exhibits greater structural diversity and frequently undergoes post-translational modifications or engages in interactions with other ..........
-
• How AI-Powered Protein Full-Length Sequencing Redefines Post-Translational Modification Analysis
The structure and function of proteins are not solely dictated by their amino acid sequences but are precisely regulated by a wide range of post-translational modifications (PTMs). Modifications such as phosphorylation, acetylation, glycosylation, and ubiquitination play pivotal roles in critical biological processes, including signal transduction, transcriptional regulation, the cell cycle, and cellular stress responses. However, the inherent diversity and complexity of PTMs pose significant challenges....
-
• Advantages and Limitations of Targeted Proteomics
As life science research transitions from “discovery” to “validation,” greater demands have been placed on the accuracy, reproducibility, and quantitative capabilities of proteomic data. Targeted Proteomics has therefore emerged as a critical bridge between basic research and clinical translation. In particular, technologies such as Multiple Reaction Monitoring (MRM) and Parallel Reaction Monitoring (PRM), known for their high controllability and reproducibility, have been widely employed in areas .........
-
In life sciences and biopharmaceutical development, the structural and functional characteristics of proteins are intrinsically linked. Comprehensive knowledge of a protein’s full-length sequence is fundamental for understanding its biological functions, identifying mutation sites, and characterizing post-translational modifications. Conventional mass spectrometry-based approaches typically rely on enzymatic digestion, fragmenting proteins into peptides for identification, which often results in partial....
-
• How to Use Targeted Proteomics for Accurate Detection of Low-Abundance Proteins
In complex biological samples, protein abundances can span a dynamic range exceeding six orders of magnitude (10⁶). Many proteins with critical biological functions—such as hormone receptors, transcription factors, signaling molecules, and early-stage disease biomarkers—fall into the category of low-abundance proteins. Despite their vital roles in various biological processes, their detection is often hindered by low expression levels and complex sample backgrounds, making conventional discovery-based......
-
• Overview of Targeted Proteomics: Methods, Techniques, Key Points, and Applications
With the increasing emphasis on precision, quantification, and clinical translation in life science research, Targeted Proteomics has emerged as a pivotal tool for advancing proteomic discoveries into clinical applications. Characterized by high sensitivity, high reproducibility, and high specificity, this approach serves as a vital link between omics-based findings and practical implementation. It is particularly well-suited for applications such as biomarker validation, pharmacodynamic studies, immune....
How to order?







