Resources
Proteomics Databases
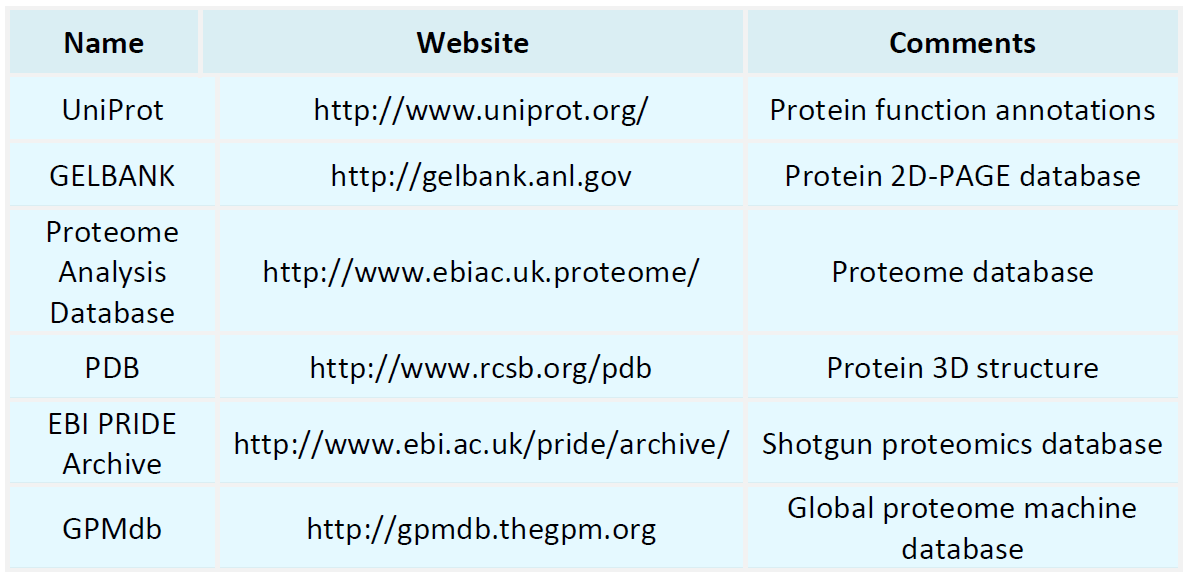
Metabolomics Databases
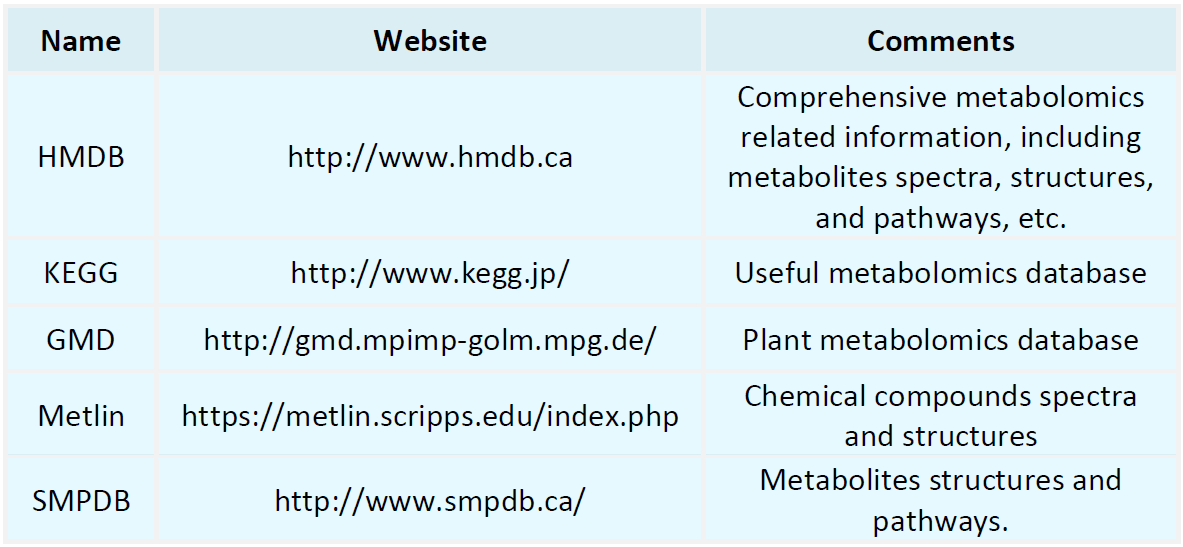
-
• Quantification of Free Thiol in Nasopharyngeal Carcinoma Tissue
In cancer research, in-depth analysis of tumor tissues is crucial. Nasopharyngeal carcinoma, as a common malignant tumor in the head and neck region, requires more precise biomarkers for diagnosis and treatment. MtoZ Biolabs is dedicated to providing important technical support for the study of nasopharyngeal carcinoma tissues using advanced free thiol quantification analysis methods.
-
Oxidation-reduction modification proteomics is an emerging scientific field that focuses on studying the impact of protein oxidation-reduction modifications on cellular function. This research area combines techniques from chemistry, biology, and proteomics to provide a new perspective on understanding cellular signal transduction, metabolic regulation, and disease mechanisms. The oxidative-reductive state of proteins directly affects their function and interactions within cells.
-
• Application of TMT and iTRAQ in Proteomics Research
TMT (Tandem Mass Tag) and iTRAQ (Isobaric Tags for Relative and Absolute Quantitation) are two commonly used mass spectrometry labeling techniques in proteomics. These techniques are used for quantitative analysis of protein expression levels in different samples and are particularly suitable for comparing the protein composition of biological samples under different conditions, such as healthy and diseased states, or different biological treatments.
-
• The Application of TMT Technology in Proteomics
The Tandem Mass Tag (TMT) quantification technology has a wide range of applications in proteomics. This technique allows researchers to simultaneously quantify proteins in multiple samples using chemically labeled tags. Here are some key examples of the applications of TMT technology: Biomarker discovery: In disease research, TMT technology is used to identify and validate potential biomarkers.
-
• Deep Quantitative Proteomics of Biological Samples via iTRAQ/TMT
iTRAQ (Isobaric Tags for Relative and Absolute Quantitation) and TMT (Tandem Mass Tag) are two commonly used mass spectrometry labeling techniques for protein quantification analysis in proteomics research. These techniques quantify the expression levels of proteins in different samples by using chemical tags, particularly suitable for comparing biological samples under different conditions, such as disease versus normal state, or cells before and after drug treatment.
-
• Proteomics of Animal Organizations
Animal tissues are composed of different types of cells, which have specific structures and functions. Animal tissues include many different types, each with specific physiological functions and characteristics. Common types of animal tissues include muscle tissue, nerve tissue, blood tissue, skin tissue, and adipose tissue, among others. Different types of animal tissues work together to maintain the normal physiological functions of the animal.
-
• Whole Blood Sample Collection in Proteomics Analysis
In proteomics analysis, the collection of whole blood samples is an important step, which is done through venous blood collection and requires the use of sterile techniques to avoid contamination. The collected blood should be immediately labeled and refrigerated to maintain sample integrity and reduce protein degradation. Additionally, the samples should be transported and stored under low temperature conditions until proteinomics analysis is performed.
-
• Precautions for Quantitative Proteomics Experiments
Quantitative proteomics experiments are important biological research methods used to measure the relative or absolute amounts of proteins in different biological samples. These experiments require strict adherence to a series of precautions to ensure the accuracy and reproducibility of the data.
-
• Quantitative Shotgun Proteomics
Quantitative shotgun proteomics is an advanced proteomic technique that allows scientists to simultaneously identify and quantify thousands of proteins in complex biological samples. Quantitative shotgun proteomics relies on the combination of high-performance liquid chromatography (HPLC) and tandem mass spectrometry (MS/MS). Proteins are enzymatically digested into peptides, which are then separated by liquid chromatography and quantitatively analyzed by mass spectrometry.
-
Protein validation aims to confirm the existence of proteins, identify their characteristics, and understand their roles and interactions in biological processes. This validation method is crucial for gaining a deeper understanding of the roles and functions of proteins in organisms and for advancing biomedical research and drug development.
How to order?







