iTRAQ/TMT/MultiNotch Quantitative Proteomics Service
Isobaric Tag for Relative Absolute Quantitation (iTRAQ) and Tandem Mass Tags (TMT) are two similar quantitative proteomic techniques developed by AB SCIEX and Thermo Fisher, respectively. The most distinctive advantage of these two techniques is that they can label up to 10 different samples and compare the relative proteomic change through LC-MS/MS analysis. Therefore, these techniques are widely applied to study proteomic changes under different treatments or in different development stages. MtoZ Biolabs is proud to offer iTRAQ/TMT analysis service to meet your research needs. On the basis of these two well-established techniques, we can also provide MultiNotch MS analysis service through MS3 or proton-transfer reactions (PTR) to further increase the accuracy of the proteomic quantitation. MtoZ Biolabs has developed a specialized platform, equipped with Q Exactive HF (Thermo Fisher), Orbitrap Fusion, Orbitrap Fusion Lumos mass spectrometers coupled with Nano-LC for iTRAQ/TMT analysis service. To better solve your research problems, we offer an all-inclusive service, including protein reduction and digestion, peptide labeling, LC-MS/MS analysis, MS data analysis and bioinformatics analysis. All you need to do is to tell us your project objective, and send us your protein samples, and we will complete all the following experiments.
The iTRAQ/TMT isobaric tags are similar and generally consist of three parts, reporter group, balance group, and reactive group. Peptides are labeled with isobaric tags by reacting with the reactive group. In MS spectrum, different iTRAQ-tag labeled peptides have the same mass, and thus appear in the same MS peaks. However, when fragmented in MS/MS, the reporter group dissociates to produce ion signals, which provide quantitative information regarding the relative amount of the peptide in the samples.
Analysis Workflow
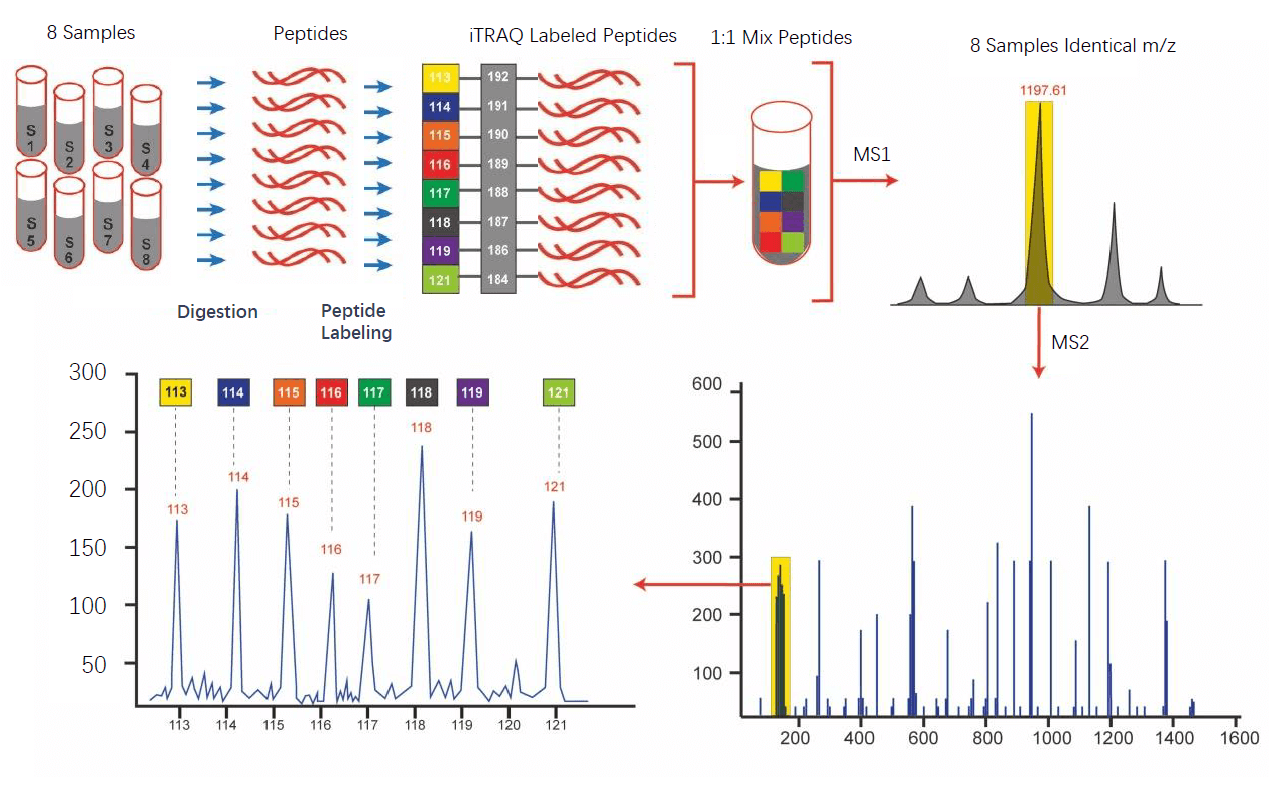
Service Advantages
1. High-Throughput: Analyze Up to 10 Samples at Once
2. In Vitro Labeling: Compatible for Labeling Various Samples, Including Tissue, Cell, Blood, etc.
3. High Sensitivity and Broad Detecting Range: Suitable for Analyzing Low Abundance Proteins
4. Compatible with Peptide Enrichment Experiment for PTMs Analysis
Applications
Analysis of large quantity of clinical samples, large scale of experimental analysis, and discovery of new biomarkers
Sample Submission Requirements
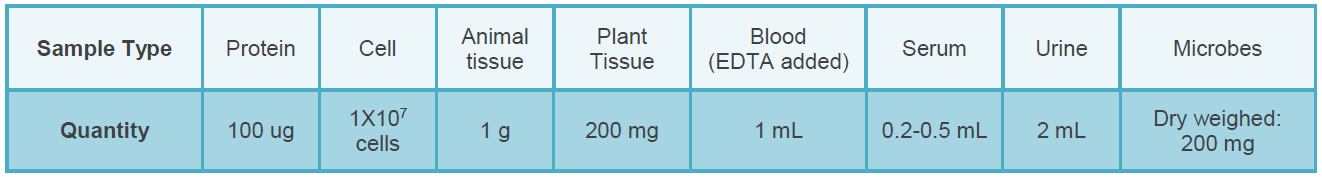
Bioinformatics Analysis
1. iTRAQ/TMT Data Quality Assessment
2. Multivariate PCA Analysis
3. Protein Statistical Analysis: Venn Diagram, Volcano Plot
4. Functional Annotation: GO Annotation, KEGG Annotation, COG Annotation
5. Clustering Analysis: Hierarchical Clustering, K-Means Clustering
6. Network Analysis: STRING Analysis
Deliverables
1. Experiment Procedures
2. Parameters of Liquid Chromatography and Mass Spectrometer
3. MS Raw Data Files
4. Peptide Identifications and Intensities
5. Protein Identifications and Intensities
6. Bioinformatics Analysis
Related Services
Quantitative Proteomics
Protein Analysis
PTM Analysis
How to order?







