N-Terminal Sequencing Service
Protein N-terminal is the start point of the expression of a protein. It also influences the subcellular distribution, degradation, and the turnover rate of a protein. Thus, sequence analysis of the N-terminal of a protein is very important for studying the function of a protein. Aimed to provide comprehensive sequencing service, MtoZ Biolabs has developed a sophisticated N-T sequencing platform consisting of Edman sequencing and protein N-T de novo sequencing, with their advantages complementary to each other.
In this service, protein sequence is not restricted by sample types, N-terminal blockage, and PTMs. Protein N-T sequences can be accurately analyzed by Edman sequencing, whereas, the N-terminal blockage and PTMs can be analyzed by mass spectrometry.
Analysis Workflow
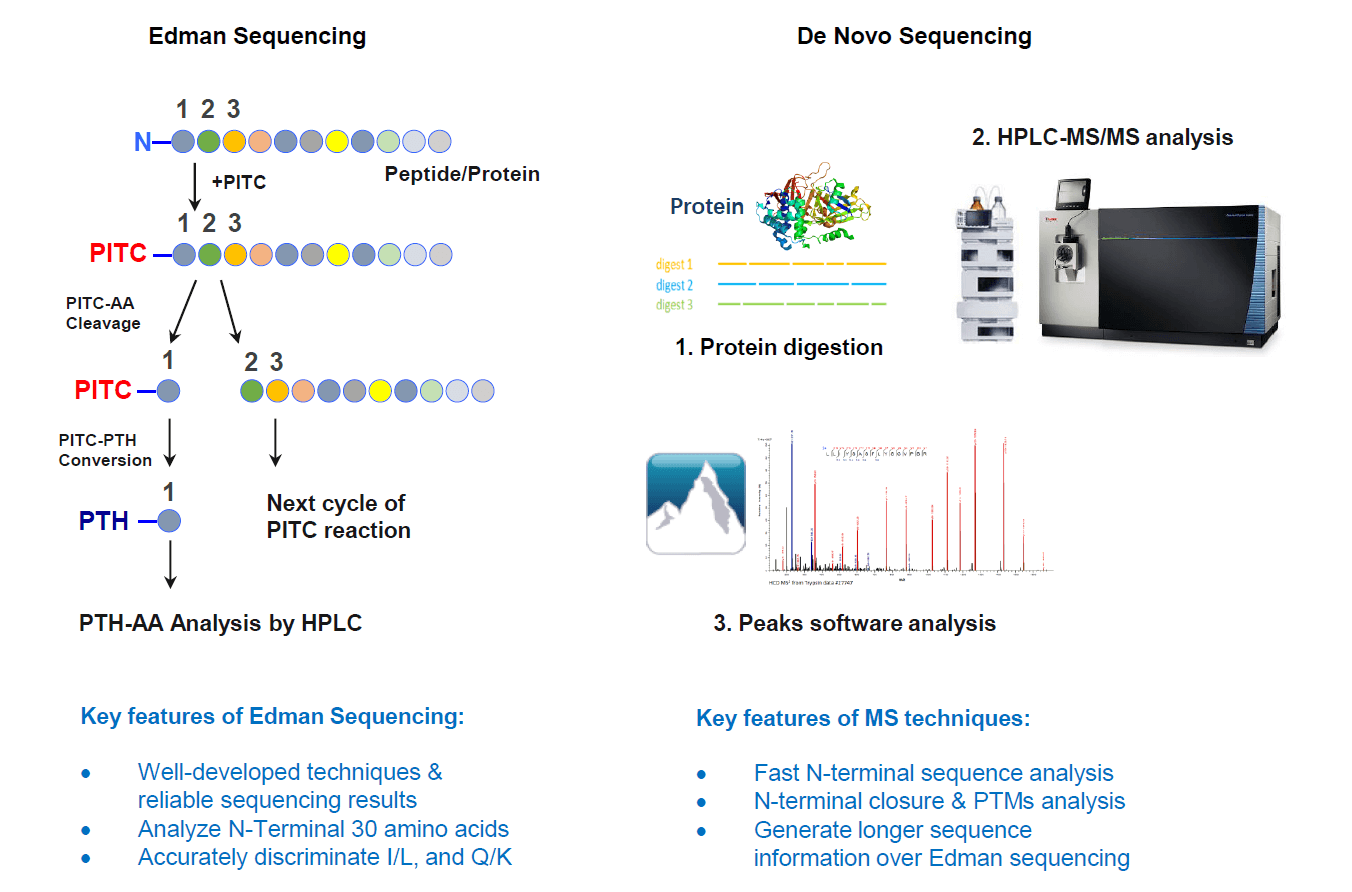
Experimental Instruments

Applications
1. Determination of N-Terminal Sequence of Proteins/Peptides/Antibodies/Vaccines
2. Analysis of PTMs
Sample Submission Requirements
1. Edman Sequencing
(1) Electroblotted Samples
Proteins separated by SDS-PAGE should be blotted onto PVDF membrane, nitrocellulose is not recommended. PVDF membrane can be stained with Coomassie blue or Poncaue red (Sliver stain is not recommended), followed by washing with ultra-pure water. The washing steps must be repeated several times, when glycine-buffer is used.
(2) Liquid Samples
1-10 ug amount of sample, with >90% purity. Avoid using Tris, glycine, guanidine, glycerol, sucrose, ethanolamine, SDS, Triton, X-100, Tween, and ammonium sulfate in the buffer.
2. MS Analysis
(1) Gel and liquid protein samples are acceptable.
(2) 5-10 ug protein samples are required. Sample purity should be as high as possible.
*Note
All reagents/solvent used must be of the highest purity to reduce contaminating substances. Samples should be handled with extreme caution and always in clean condition. Any source that may introduce contaminating proteins should be eliminated. Customers are welcome to contact us for detailed sample requirements before sending your samples.
Deliverables
1. Experiment Procedures
2. Parameters of Liquid Chromatography and Mass Spectrometer
3. MS Raw Data Files
4. Peptide Identifications and Intensity
5. Protein Identifications and Intensity
Related Services
Protein Sequencing
Protein Full-Length Sequencing
Protein Analysis
PTM Analysis
How to order?







